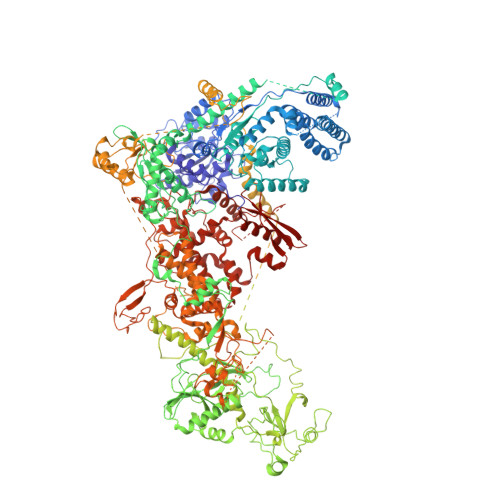
Structural basis of microRNA biogenesis by Dicer-1 and its partner protein Loqs-PB.
Jouravleva, K., Golovenko, D., Demo, G., Dutcher, R.C., Hall, T.M.T., Zamore, P.D., Korostelev, A.A.(2022) Mol Cell 82: 4049-4063.e6
- PubMed: 36182693 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2022.09.002
- Primary Citation Related Structures:
8DFV, 8DG5, 8DG7, 8DGA, 8DGI, 8DGJ - PubMed Abstract:
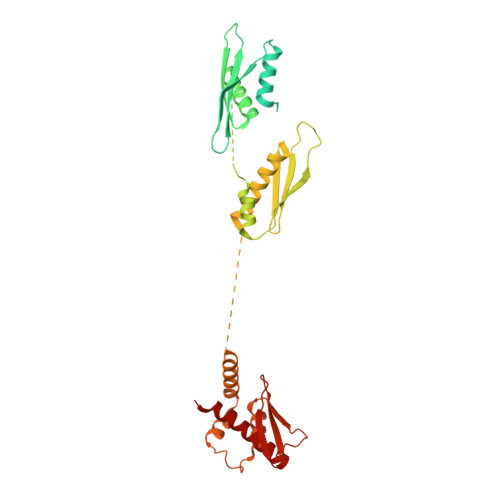
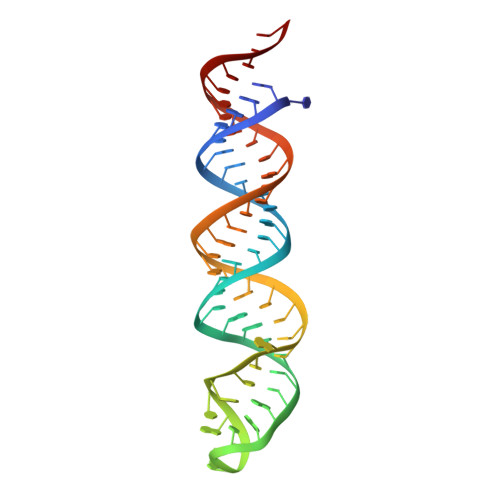
In animals and plants, Dicer enzymes collaborate with double-stranded RNA-binding domain (dsRBD) proteins to convert precursor-microRNAs (pre-miRNAs) into miRNA duplexes. We report six cryo-EM structures of Drosophila Dicer-1 that show how Dicer-1 and its partner Loqs‑PB cooperate (1) before binding pre-miRNA, (2) after binding and in a catalytically competent state, (3) after nicking one arm of the pre-miRNA, and (4) following complete dicing and initial product release. Our reconstructions suggest that pre-miRNA binds a rare, open conformation of the Dicer‑1⋅Loqs‑PB heterodimer. The Dicer-1 dsRBD and three Loqs‑PB dsRBDs form a tight belt around the pre-miRNA, distorting the RNA helix to place the scissile phosphodiester bonds in the RNase III active sites. Pre-miRNA cleavage shifts the dsRBDs and partially closes Dicer-1, which may promote product release. Our data suggest a model for how the Dicer‑1⋅Loqs‑PB complex affects a complete cycle of pre-miRNA recognition, stepwise endonuclease cleavage, and product release.
- RNA Therapeutics Institute, University of Massachusetts Chan Medical School, 368 Plantation Street, Worcester, MA 01605, USA.
Organizational Affiliation: