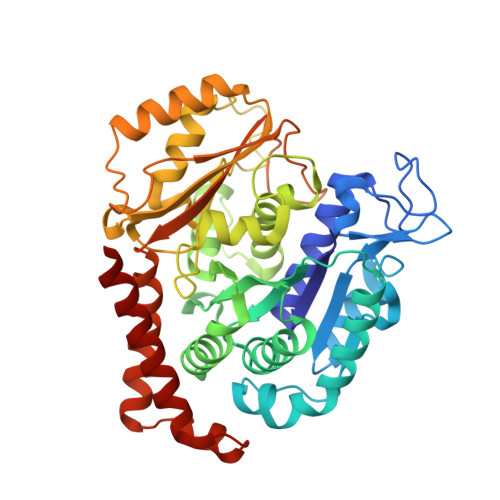
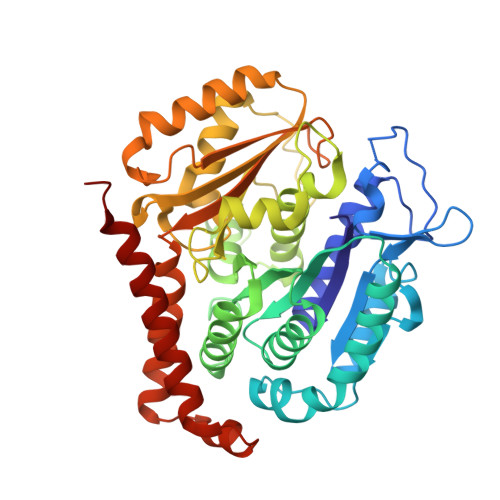
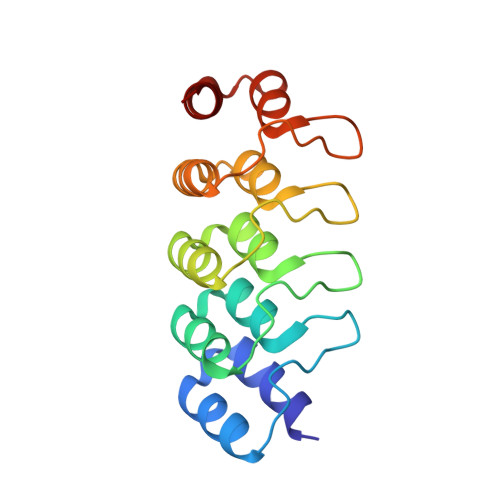
A multi-reservoir extruder for time-resolved serial protein crystallography and compound screening at X-ray free electron lasers
Wranik, M., Kepa, M.W., Beale, E.V., James, D., Bertrand, Q., Weinert, T., Furrer, A., Glover, H., Gashi, D., Carrillo, M., Kondo, Y., Stipp, R.T., Khusainov, G., Nass, K., Ozerov, D., Cirelli, C., Johnson, P.J.M., Dworkowski, F., Beale, J.H., Stubbs, S., Schneider, M., Krauskopf, K., Gao, L., Thorn-Seshold, O., Bostedt, C., Bacellar, C., Steinmetz, M.O., Milne, C., Standfuss, J.To be published.