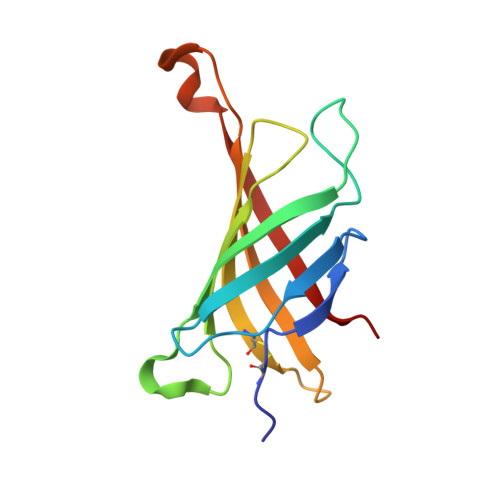
The avidin-theophylline complex: A structural and computational study.
Spinello, A., Lapenta, F., De March, M.(2023) Proteins 91: 1437-1443
- PubMed: 37318226 Search on PubMed
- DOI: https://doi.org/10.1002/prot.26538
- Primary Citation Related Structures:
8CK7 - PubMed Abstract:
The interaction between avidin and its counterpart biotin is one of central importance in biology and has been reproposed and studied at length. However, the binding pocket of avidin is prone to promiscuous binding, able to accommodate even non-biotinylated ligands. Comprehending the factors that distinguish the extremely strong interaction with biotin to other ligands is an important step to fully picture the thermodynamics of these low-affinity complexes. Here, we present the complex between chicken white egg avidin and theophylline (TEP), the xanthine derivative used in the therapy of asthma. In the crystal structure, TEP lies in the biotin-binding pocket with the same orientation and planarity of the aromatic ring of 8-oxodeoxyguanosine. Indeed, its affinity for avidin measured by isothermal titration calorimetry is in the same μM range as those obtained for the previously characterized nucleoside derivatives. By the use of molecular dynamic simulations, we have investigated the most important intermolecular interactions occurring in the avidin-TEP binding pocket and compared them with those obtained for the avidin 8-oxodeoxyguanosine and avidin-biotin complexes. These results testify the capability of avidin to complex purely aromatic molecules.
- Department of Biological, Chemical and Pharmaceutical Sciences and Technologies, University of Palermo, Palermo, Italy.
Organizational Affiliation: