CryoEM structure of ACP crosslinked to KS of the cercosporin fungal non-reducing polyketide synthase (NR-PKS) CTB1 (SAT-KS:ACP-MAT) at 2.7 Angstroms resolution
Munoz-Hernandez, H., Maier, T.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Non-reducing polyketide synthase CTB1 | 1,304 | Cercospora nicotianae | Mutation(s): 3 Gene Names: CTB1 EC: 2.3.1 | 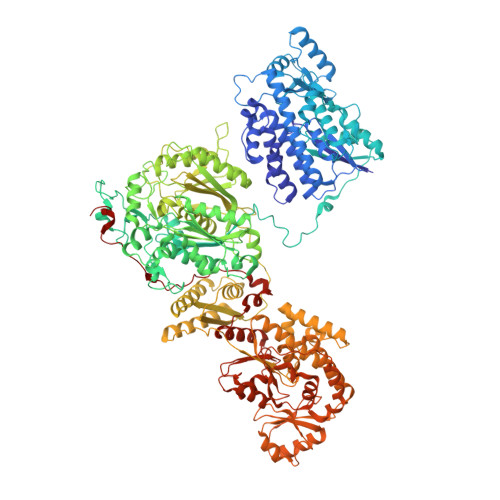 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q6DQW3 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Acyl carrier protein (ACP) of Non-reducing polyketide synthase CTB1 | 88 | Cercospora nicotianae | Mutation(s): 0 Gene Names: CTB1 EC: 2.3.1 | 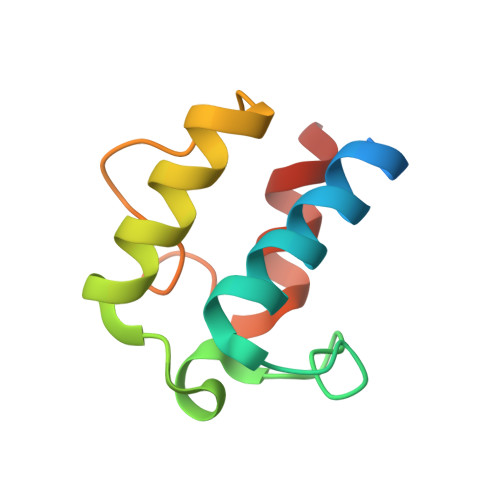 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q6DQW3 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| 42X (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | D [auth C] | N~3~-[(2R)-2-hydroxy-3,3-dimethyl-4-(phosphonooxy)butanoyl]-N-[2-(propanoylamino)ethyl]-beta-alaninamide C14 H28 N3 O8 P IEZRALNUCGHOKF-LBPRGKRZSA-N |  | ||
| Task | Software Package | Version |
|---|---|---|
| RECONSTRUCTION | cryoSPARC | |
| MODEL REFINEMENT | PHENIX |
| Funding Organization | Location | Grant Number |
|---|---|---|
| European Commission | European Union | ID: 845941 |
| Swiss National Science Foundation | Switzerland | -- |