Anti-GARP Antibodies Inhibit Release of TGF-beta by Regulatory T Cells via Different Modes of Action, but Do Not Influence Their Function In Vitro.
Igney, F.H., Ebenhoch, R., Schiele, F., Nar, H.(2023) Immunohorizons 7: 200-212
- PubMed: 36928178 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.4049/immunohorizons.2200072
- Primary Citation Related Structures:
8C7H - PubMed Abstract:
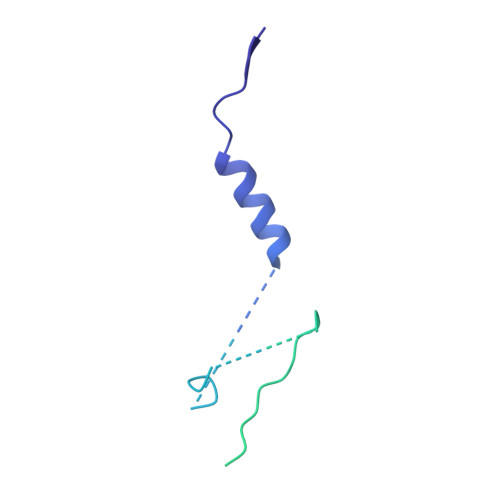
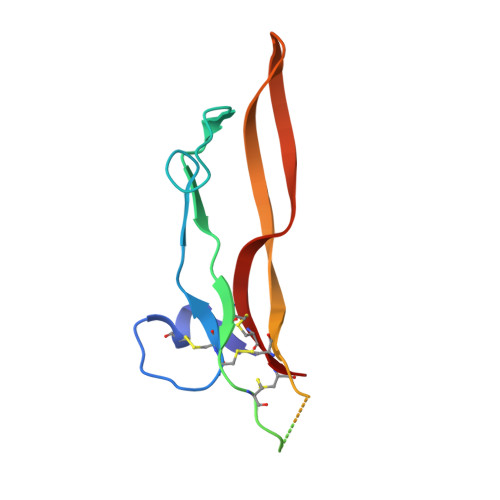
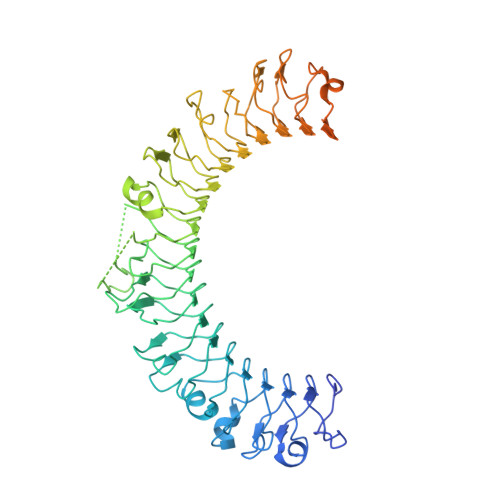
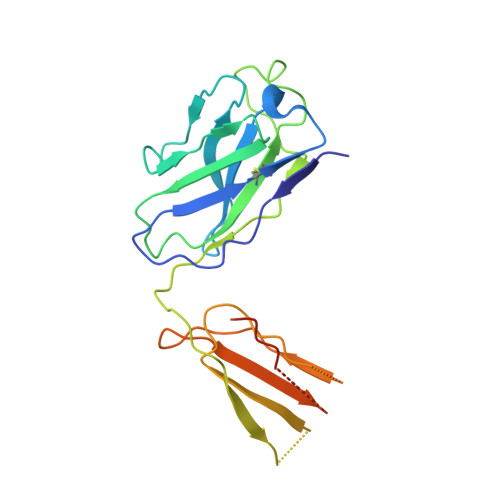
Regulatory T cells (Treg) play a critical role in controlling immune responses in diseases such as cancer or autoimmunity. Activated Treg express the membrane protein GARP (LRRC32) in complex with the latent form of the immunosuppressive cytokine TGF-β (L-TGF-β). In this study, we confirmed that active TGF-β was generated from its latent form in an integrin-dependent manner and induced TGF-β receptor signaling in activated human Treg. We studied a series of Abs targeting the L-TGF-β/GARP complex with distinct binding modes. We found that TGF-β receptor signaling could be inhibited by anti-TGF-β and by some, but not all, Abs against the L-TGF-β/GARP complex. Cryogenic electron microscopy structures of three L-TGF-β/GARP complex-targeting Abs revealed their distinct epitopes and allowed us to elucidate how they achieve blockade of TGF-β activation. Three different modes of action were identified, including a novel unusual mechanism of a GARP-binding Ab. However, blockade of GARP or TGF-β by Abs did not influence the suppressive activity of human Treg in vitro. We were also not able to confirm a prominent role of GARP in other functions of human Treg, such as FOXP3 induction and Treg stability. These data show that the GARP/TGF-β axis can be targeted pharmacologically in different ways, but further studies are necessary to understand its complexity and to unleash its therapeutic potential.
- Discovery Research, Cancer Immunology & Immune Modulation, Boehringer Ingelheim Pharma GmbH & Co. KG, Biberach an der Riss, Germany.
Organizational Affiliation: