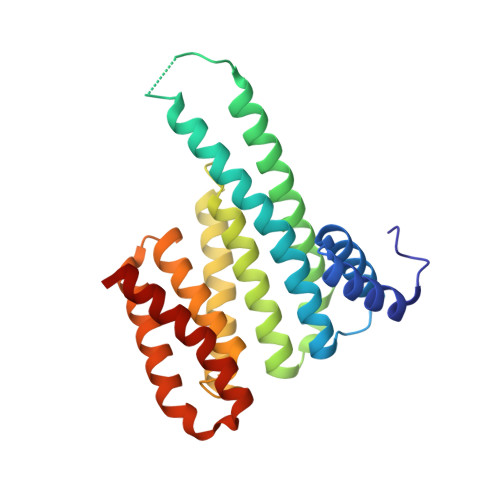
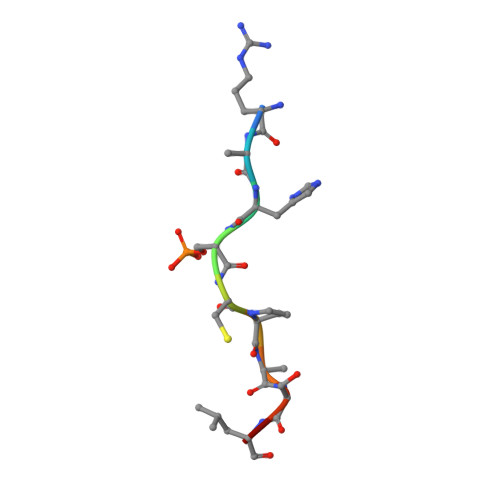
A simple method for developing lysine targeted covalent protein reagents.
Gabizon, R., Tivon, B., Reddi, R.N., van den Oetelaar, M.C.M., Amartely, H., Cossar, P.J., Ottmann, C., London, N.(2023) Nat Commun 14: 7933-7933
- PubMed: 38040731 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-42632-5
- Primary Citation Related Structures:
8C2E, 8C2F - PubMed Abstract:
Peptide-based covalent probes can target shallow protein surfaces not typically addressable using small molecules, yet there is a need for versatile approaches to convert native peptide sequences into covalent binders that can target a broad range of residues. Here we report protein-based thio-methacrylate esters-electrophiles that can be installed easily on unprotected peptides and proteins via cysteine side chains, and react efficiently and selectively with cysteine and lysine side chains on the target. Methacrylate phosphopeptides derived from 14-3-3-binding proteins irreversibly label 14-3-3σ via either lysine or cysteine residues, depending on the position of the electrophile. Methacrylate peptides targeting a conserved lysine residue exhibit pan-isoform binding of 14-3-3 proteins both in lysates and in extracellular media. Finally, we apply this approach to develop protein-based covalent binders. A methacrylate-modified variant of the colicin E9 immunity protein irreversibly binds to the E9 DNAse, resulting in significantly higher thermal stability relative to the non-covalent complex. Our approach offers a simple and versatile route to convert peptides and proteins into potent covalent binders.
- Department of Chemical and Structural Biology, The Weizmann Institute of Science, Rehovot, 7610001, Israel.
Organizational Affiliation: