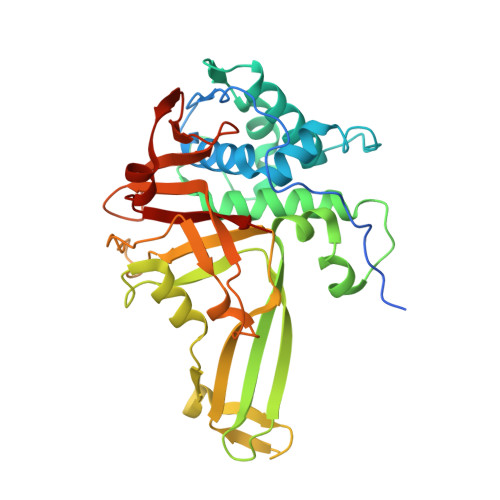
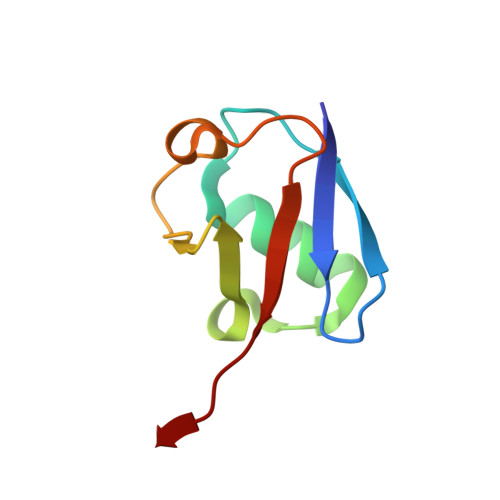
Molecular basis for ubiquitin/Fubi cross-reactivity in USP16 and USP36.
O'Dea, R., Kazi, N., Hoffmann-Benito, A., Zhao, Z., Recknagel, S., Wendrich, K., Janning, P., Gersch, M.(2023) Nat Chem Biol 19: 1394-1405
- PubMed: 37443395 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-023-01388-1
- Primary Citation Related Structures:
8BS3, 8BS9 - PubMed Abstract:
Ubiquitin and ubiquitin-like proteins typically use distinct machineries to facilitate diverse functions. The immunosuppressive ubiquitin-like protein Fubi is synthesized as an N-terminal fusion to a ribosomal protein (Fubi-S30). Its proteolytic maturation by the nucleolar deubiquitinase USP36 is strictly required for translationally competent ribosomes. What endows USP36 with this activity, how Fubi is recognized and whether other Fubi proteases exist are unclear. Here, we report a chemical tool kit that facilitated the discovery of dual ubiquitin/Fubi cleavage activity in USP16 in addition to USP36 by chemoproteomics. Crystal structures of USP36 complexed with Fubi and ubiquitin uncover its substrate recognition mechanism and explain how other deubiquitinases are restricted from Fubi. Furthermore, we introduce Fubi C-terminal hydrolase measurements and reveal a synergistic role of USP16 in Fubi-S30 maturation. Our data highlight how ubiquitin/Fubi specificity is achieved in a subset of human deubiquitinases and open the door to a systematic investigation of the Fubi system.
- Chemical Genomics Centre, Max Planck Institute of Molecular Physiology, Dortmund, Germany.
Organizational Affiliation: