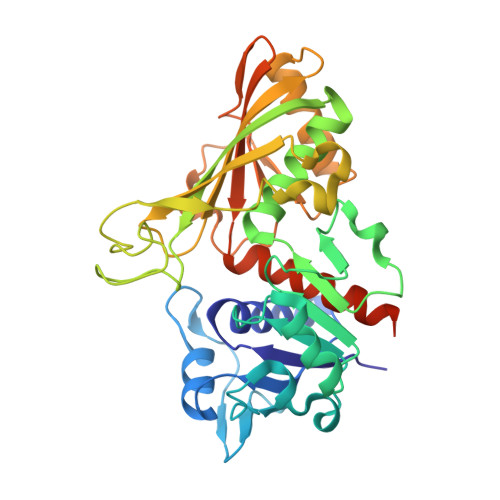
Stoichiometry of ligand binding and role of C-terminal lysines in Mycobacterium tuberculosis and human GAPDH multifunctionality.
Kumar, A., Kumar, R., Boradia, V.M., Malhotra, H., Kumar, A., Seth, S., Garg, P., Karthikeyan, S., Raje, M., Iyengar Raje, C.(2024) FEBS J
- PubMed: 39436721 Search on PubMed
- DOI: https://doi.org/10.1111/febs.17298
- Primary Citation Related Structures:
8BR4 - PubMed Abstract:
Glyceraldehyde-3-phosphate-dehydrogenase (GAPDH; EC1.2.1.12) has several functions in Mycobacterium tuberculosis (Mtb) and the human host. Apart from its role in glycolysis, it serves both as a cell surface and a secreted receptor for plasmin(ogen) (Plg/Plm), transferrin (Tf), and lactoferrin (Lf). Plg sequestration by Mtb GAPDH facilitates bacterial adhesion and tissue invasion, while an equivalent interaction with host GAPDH regulates immune cell migration. In both, host and microbe, internalization of Tf/Lf-GAPDH complexes serves as a route for iron acquisition. To date, the structure of Mtb GAPDH or the residues involved in these moonlighting interactions have not been identified. This study provides the first known X-ray crystal structure of Mtb GAPDH. Through further mutagenesis and functional assays, we found that the C-terminal lysines of Mtb and human GAPDH affect enzyme activity and ligand binding. We also establish the stoichiometry of Plg, Tf and Lf interactions with the GAPDH tetramer. Lastly, molecular simulation studies reveal the interactions of the C-terminal lysine residues.
- Department of Biotechnology, National Institute of Pharmaceutical Education and Research, Sahibzada Ajit Singh Nagar, Punjab, India.
Organizational Affiliation: