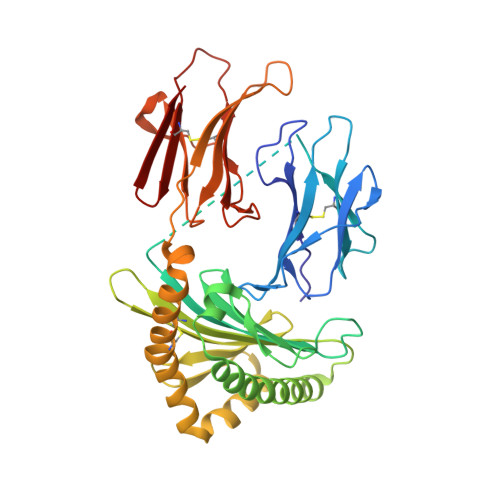
The carbonyl nucleobase adduct M 3 Ade is a potent antigen for adaptive polyclonal MR1-restricted T cells.
Chancellor, A., Constantin, D., Berloffa, G., Yang, Q., Nosi, V., Loureiro, J.P., Colombo, R., Jakob, R.P., Joss, D., Pfeffer, M., De Simone, G., Morabito, A., Schaefer, V., Vacchini, A., Brunelli, L., Montagna, D., Heim, M., Zippelius, A., Davoli, E., Haussinger, D., Maier, T., Mori, L., De Libero, G.(2024) Immunity
- PubMed: 39701104 Search on PubMed
- DOI: https://doi.org/10.1016/j.immuni.2024.11.019
- Primary Citation Related Structures:
8BP6 - PubMed Abstract:
The major histocompatibility complex (MHC) class I-related molecule MHC-class-I-related protein 1 (MR1) presents metabolites to distinct MR1-restricted T cell subsets, including mucosal-associated invariant T (MAIT) and MR1T cells. However, self-reactive MR1T cells and the nature of recognized antigens remain underexplored. Here, we report a cell endogenous carbonyl adduct of adenine (8-(9H-purin-6-yl)-2-oxa-8-azabicyclo[3.3.1]nona-3,6-diene-4,6-dicarbaldehyde [M 3 Ade]) sequestered in the A' pocket of MR1. M 3 Ade induced in vitro MR1-mediated stimulation of MR1T cell clones that bound MR1-M 3 Ade tetramers. MR1-M 3 Ade tetramers identified heterogeneous MR1-reactive T cells ex vivo in healthy donors, individuals with acute myeloid leukemia, and tumor-infiltrating lymphocytes from non-small cell lung adenocarcinoma and hepatocarcinoma. These cells displayed phenotypic, transcriptional, and functional diversity at distinct differentiation stages, indicating their adaptive nature. They were also polyclonal, with some preferential T cell receptor (TCRαβ) pair usage. Thus, M 3 Ade is an MR1-presented self-metabolite that enables stimulation and tracking of human-MR1T cells from blood and tissue, aiding our understanding of their roles in health and disease.
- Experimental Immunology, Department of Biomedicine, University Hospital Basel, University of Basel, 4031 Basel, Switzerland. Electronic address: andrew.chancellor@unibas.ch.
Organizational Affiliation: