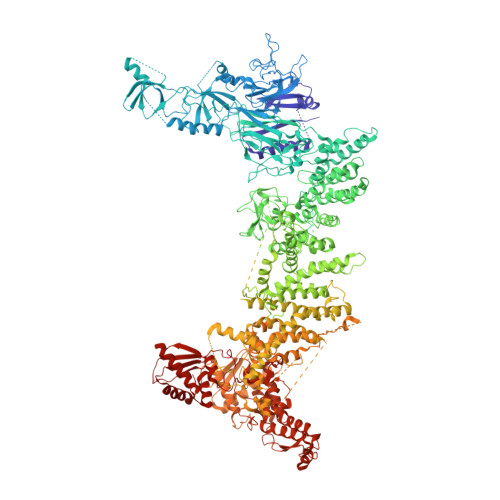
Cryo-EM structure of the chain-elongating E3 ubiquitin ligase UBR5.
Hodakova, Z., Grishkovskaya, I., Brunner, H.L., Bolhuis, D.L., Belacic, K., Schleiffer, A., Kotisch, H., Brown, N.G., Haselbach, D.(2023) EMBO J 42: e113348-e113348
- PubMed: 37409633 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embj.2022113348
- Primary Citation Related Structures:
8BJA - PubMed Abstract:
UBR5 is a nuclear E3 ligase that ubiquitinates a vast range of substrates for proteasomal degradation. This HECT domain-containing ubiquitin ligase has recently been identified as an important regulator of oncogenes, e.g., MYC, but little is known about its structure or mechanisms of substrate engagement and ubiquitination. Here, we present the cryo-EM structure of human UBR5, revealing an α-solenoid scaffold with numerous protein-protein interacting motifs, assembled into an antiparallel dimer that adopts further oligomeric states. Using cryo-EM processing tools, we observe the dynamic nature of the UBR5 catalytic domain, which we postulate is important for its enzymatic activity. We characterise the proteasomal nuclear import factor AKIRIN2 as an interacting protein and propose UBR5 as an efficient ubiquitin chain elongator. This preference for ubiquitinated substrates and several distinct domains for protein-protein interactions may explain how UBR5 is linked to several different signalling pathways and cancers. Together, our data expand on the limited knowledge of the structure and function of HECT E3 ligases.
- Research Institute of Molecular Pathology (IMP), ViennaBioCenter (VBC), Vienna, Austria.
Organizational Affiliation: