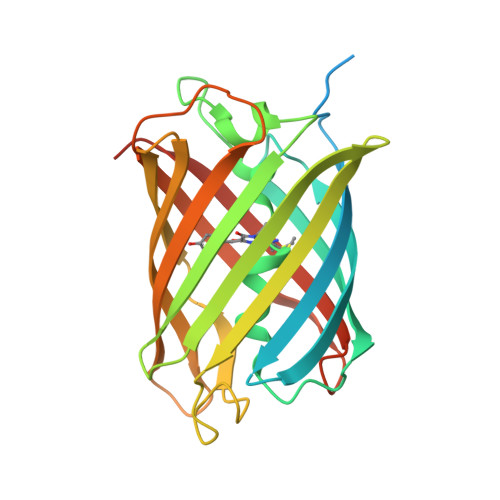
An unusual disulfide-linked dimerization in the fluorescent protein rsCherryRev1.4.
Bui, T.Y.H., Dedecker, P., Van Meervelt, L.(2023) Acta Crystallogr F Struct Biol Commun 79: 38-44
- PubMed: 36748340 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X23000572
- Primary Citation Related Structures:
8BGL - PubMed Abstract:
rsCherryRev1.4 has been reported as one of the reversibly photoswitchable variants of mCherry, and is an improved version with a faster off-switching speed and lower switching fatigue at high light intensities than its precursor rsCherryRev. However, rsCherryRev1.4 still has some limitations such as a tendency to dimerize as well as complex photophysical properties. Here, the crystal structure of rsCherryRev1.4 was determined at a resolution of 2 Å and it was discovered that it forms a dimer that shows disulfide bonding between the protomers. Mutagenesis, gel electrophoresis and size-exclusion chromatography strongly implicate Cys24 in this process. Replacing Cys24 in rsCherryRev1.4 resulted in a much lower tendency towards dimerization, while introducing Cys24 into mCherry correspondingly increased its dimerization. In principle, this finding opens the possibility of developing redox sensors based on controlled dimerization via disulfide cross-linking in fluorescent proteins, even though the actual application of engineering such sensors still requires additional research.
- Biochemistry, Molecular and Structural Biology, Department of Chemistry, KU Leuven, 3001 Leuven, Belgium.
Organizational Affiliation: