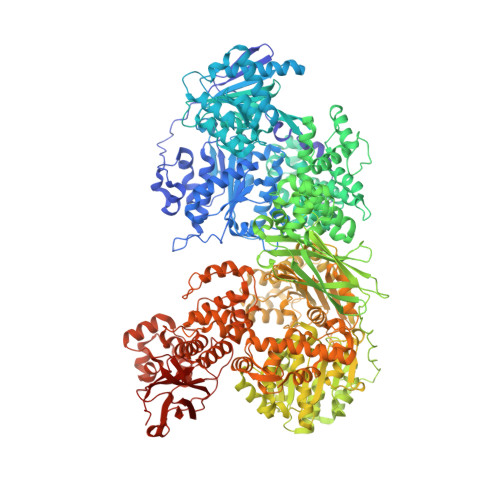
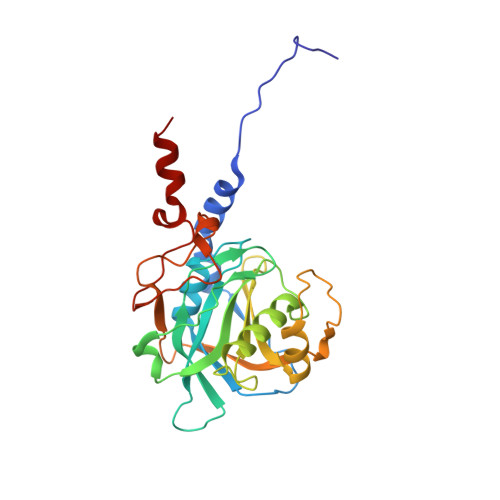
Conformation-dependent ligand hot spots in the spliceosomal RNA helicase BRR2.
Vester, K., Metz, A., Huber, S., Loll, B., Wahl, M.C.(2023) Acta Crystallogr D Struct Biol 79: 304-317
- PubMed: 36974964 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798323001778
- Primary Citation Related Structures:
8BC8, 8BC9, 8BCA, 8BCB, 8BCC, 8BCD, 8BCE, 8BCF, 8BCG, 8BCH - PubMed Abstract:
The conversion of hits to leads in drug discovery involves the elaboration of chemical core structures to increase their potency. In fragment-based drug discovery, low-molecular-weight compounds are tested for protein binding and are subsequently modified, with the tacit assumption that the binding mode of the original hit will be conserved among the derivatives. However, deviations from binding mode conservation are rather frequently observed, but potential causes of these alterations remain incompletely understood. Here, two crystal forms of the spliceosomal RNA helicase BRR2 were employed as a test case to explore the consequences of conformational changes in the target protein on the binding behaviour of fragment derivatives. The initial fragment, sulfaguanidine, bound at the interface between the two helicase cassettes of BRR2 in one crystal form. Second-generation compounds devised by structure-guided docking were probed for their binding to BRR2 in a second crystal form, in which the original fragment-binding site was altered due to a conformational change. While some of the second-generation compounds retained binding to parts of the original site, others changed to different binding pockets of the protein. A structural bioinformatics analysis revealed that the fragment-binding sites correspond to predicted binding hot spots, which strongly depend on the protein conformation. This case study offers an example of extensive binding-mode changes during hit derivatization, which are likely to occur as a consequence of multiple binding hot spots, some of which are sensitive to the flexibility of the protein.
- Laboratory of Structural Biochemistry, Institute of Chemistry and Biochemistry, Freie Universität Berlin, Takustrasse 6, 14195 Berlin, Germany.
Organizational Affiliation: