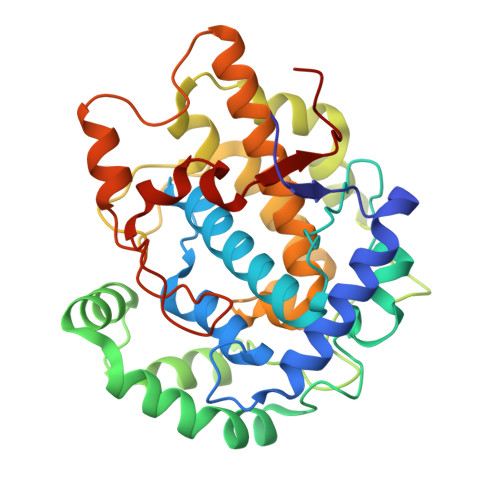
The Crystal Structure of Tyrosinase from Verrucomicrobium spinosum Reveals It to Be an Atypical Bacterial Tyrosinase.
Fekry, M., Dave, K.K., Badgujar, D., Hamnevik, E., Aurelius, O., Dobritzsch, D., Danielson, U.H.(2023) Biomolecules 13
- PubMed: 37759761 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/biom13091360
- Primary Citation Related Structures:
8BBQ, 8BBR - PubMed Abstract:
Tyrosinases belong to the type-III copper enzyme family, which is involved in melanin production in a wide range of organisms. Despite similar overall characteristics and functions, their structures, activities, substrate specificities and regulation vary. The tyrosinase from the bacterium Verrucomicrobium spinosum ( vs Tyr) is produced as a pre-pro-enzyme in which a C-terminal extension serves as an inactivation domain. It does not require a caddie protein for copper ion incorporation, which makes it similar to eukaryotic tyrosinases. To gain an understanding of the catalytic machinery and regulation of vs Tyr activity, we determined the structure of the catalytically active "core domain" of vs Tyr by X-ray crystallography. The analysis showed that vs Tyr is an atypical bacterial tyrosinase not only because it is independent of a caddie protein but also because it shows the highest structural (and sequence) similarity to plant-derived members of the type-III copper enzyme family and is more closely related to fungal tyrosinases regarding active site features. By modelling the structure of the pre-pro-enzyme using AlphaFold, we observed that Phe453, located in the C-terminal extension, is appropriately positioned to function as a "gatekeeper" residue. Our findings raise questions concerning the evolutionary origin of vs Tyr.
- Department of Chemistry-BMC, Uppsala University, SE 751 23 Uppsala, Sweden.
Organizational Affiliation: