Nanobody inhibitors of Plexin-B1 identify allostery in plexin-semaphorin interactions and signaling.
Cowan, R., Trokter, M., Oleksy, A., Fedorova, M., Sawmynaden, K., Worzfeld, T., Offermanns, S., Matthews, D., Carr, M.D., Hall, G.(2023) J Biological Chem 299: 104740-104740
- PubMed: 37088134 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2023.104740
- Primary Citation Related Structures:
8B3K, 8BB7, 8BF4 - PubMed Abstract:
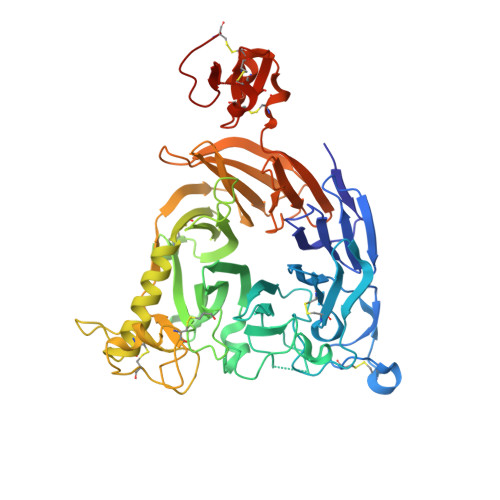
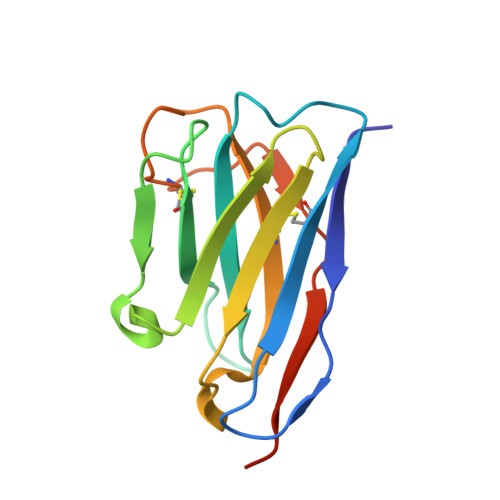
Plexin-B1 is a receptor for the cell surface semaphorin, Sema4D. This signaling system has been implicated in a variety of human diseases, including cancer, multiple sclerosis and osteoporosis. While inhibitors of the Plexin-B1:Sema4D interaction have been previously reported, understanding their mechanism has been hindered by an incomplete structural view of Plexin-B1. In this study, we have raised and characterized a pair of nanobodies that are specific for mouse Plexin-B1 and which inhibit the binding of Sema4D to mouse Plexin-B1 and its biological activity. Structural studies of these nanobodies reveal that they inhibit the binding of Sema4D in an allosteric manner, binding to epitopes not previously reported. In addition, we report the first unbound structure of human Plexin-B1, which reveals that Plexin-B1 undergoes a conformational change on Sema4D binding. These changes mirror those seen upon binding of allosteric peptide modulators, which suggests a new model for understanding Plexin-B1 signaling and provides a potential innovative route for therapeutic modulation of Plexin-B1.
- Department of Molecular and Cell Biology, Leicester Institute of Structural and Chemical Biology, University of Leicester, Leicester, UK.
Organizational Affiliation: