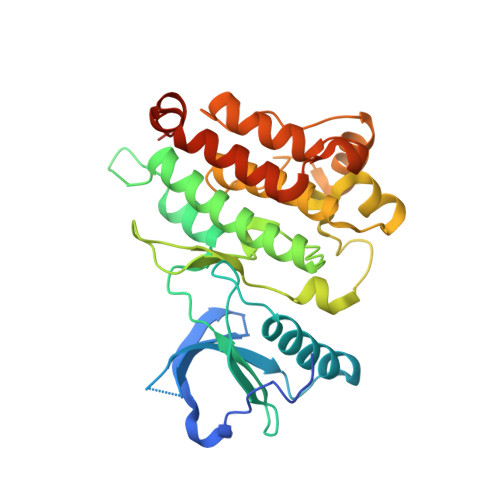
Discovery of Novel alpha-Carboline Inhibitors of the Anaplastic Lymphoma Kinase.
Mologni, L., Tardy, S., Zambon, A., Orsato, A., Bisson, W.H., Ceccon, M., Viltadi, M., D'Attoma, J., Pannilunghi, S., Vece, V., Bertho, J., Goekjian, P., Scapozza, L., Gambacorti-Passerini, C.(2022) ACS Omega 7: 17083-17097
- PubMed: 35647450 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsomega.2c00507
- Primary Citation Related Structures:
8ARJ - PubMed Abstract:
The anaplastic lymphoma kinase (ALK) is abnormally expressed and hyperactivated in a number of tumors and represents an ideal therapeutic target. Despite excellent clinical responses to ALK inhibition, drug resistance still represents an issue and novel compounds that overcome drug-resistant mutants are needed. We designed, synthesized, and evaluated a large series of azacarbazole inhibitors. Several lead compounds endowed with submicromolar potency were identified. Compound 149 showed selective inhibition of native and mutant drug-refractory ALK kinase in vitro as well as in a Ba/F3 model and in human ALK+ lymphoma cells. The three-dimensional (3D) structure of a 149 :ALK-KD cocrystal is reported, showing extensive interaction through the hinge region and the catalytic lysine 1150.
- Dept. of Medicine and Surgery, University of Milano-Bicocca, Monza 20900, Italy.
Organizational Affiliation: