Identification, binding, and structural characterization of single domain anti-PD-L1 antibodies inhibitory of immune regulatory proteins PD-1 and CD80.
Kang-Pettinger, T., Walker, K., Brown, R., Cowan, R., Wright, H., Baravalle, R., Waters, L.C., Muskett, F.W., Bowler, M.W., Sawmynaden, K., Coombs, P.J., Carr, M.D., Hall, G.(2023) J Biological Chem 299: 102769-102769
- PubMed: 36470427 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2022.102769
- Primary Citation Related Structures:
8AOK, 8AOM - PubMed Abstract:
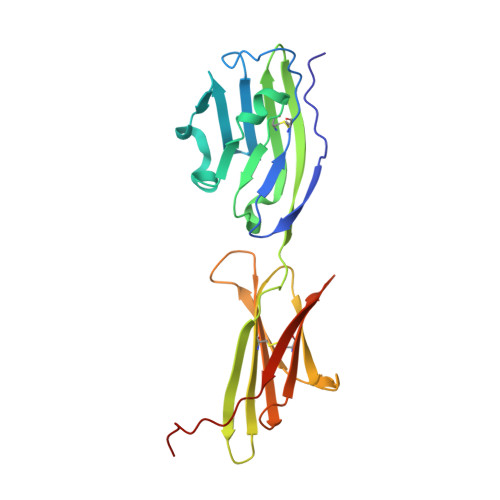
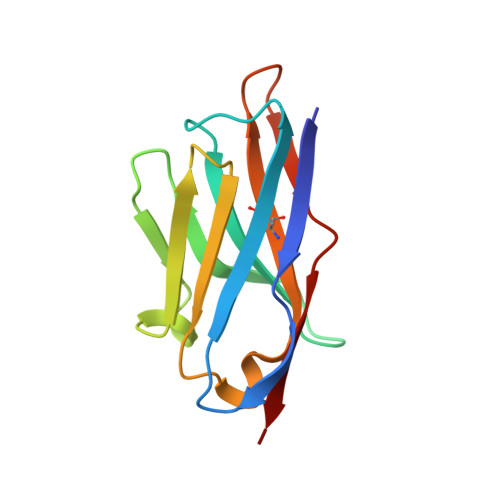
Programmed death-ligand 1 (PD-L1) is a key immune regulatory protein that interacts with programmed cell death protein 1 (PD-1), leading to T-cell suppression. Whilst this interaction is key in self-tolerance, cancer cells evade the immune system by overexpressing PD-L1. Inhibition of the PD-1/PD-L1 pathway with standard monoclonal antibodies has proven a highly effective cancer treatment; however, single domain antibodies (VHH) may offer numerous potential benefits. Here, we report the identification and characterization of a diverse panel of 16 novel VHHs specific to PD-L1. The panel of VHHs demonstrate affinities of 0.7 nM to 5.1 μM and were able to completely inhibit PD-1 binding to PD-L1. The binding site for each VHH on PD-L1 was determined using NMR chemical shift perturbation mapping and revealed a common binding surface encompassing the PD-1-binding site. Additionally, we solved crystal structures of two representative VHHs in complex with PD-L1, which revealed unique binding modes. Similar NMR experiments were used to identify the binding site of CD80 on PD-L1, which is another immune response regulatory element and interacts with PD-L1 localized on the same cell surface. CD80 and PD-1 were revealed to share a highly overlapping binding site on PD-L1, with the panel of VHHs identified expected to inhibit CD80 binding. Comparison of the CD80 and PD-1 binding sites on PD-L1 enabled the identification of a potential antibody binding region able to confer specificity for the inhibition of PD-1 binding only, which may offer therapeutic benefits to counteract cancer cell evasion of the immune system.
- Leicester Institute of Structural and Chemical Biology and Department of Molecular and Cell Biology, Henry Wellcome Building, University of Leicester, Leicester, UK.
Organizational Affiliation: