3D Structure of D-Аmino Acid Тransaminase from Aminobacterium colombiense in Complex with D-Cycloserine
Shilova, S.A., Matyuta, I.O., Bezsudnova, E.Y., Minyaev, M.E., Nikolaeva, A.Y., Popov, V.O., Boyko, K.M.(2023) Crystallogr Rep 68: 931-937
Experimental Data Snapshot
Starting Model: experimental
View more details
(2023) Crystallogr Rep 68: 931-937
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Aminotransferase class IV | 277 | Aminobacterium colombiense | Mutation(s): 0 Gene Names: Amico_1844 | 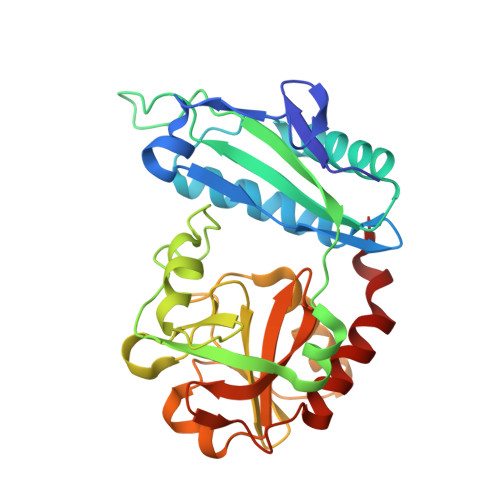 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | D5EHC5 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| M7L (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | E [auth B] | 3-azanyloxy-2-[(~{E})-[2-methyl-3-oxidanyl-5-(phosphonooxymethyl)pyridin-4-yl]methylideneamino]propanoic acid C11 H16 N3 O8 P ICHYGZXBYNDCFX-NTEUORMPSA-N |  | ||
| LCS (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | C [auth A] | [5-hydroxy-6-methyl-4-({[(4E)-3-oxo-1,2-oxazolidin-4-ylidene]amino}methyl)pyridin-3-yl]methyl dihydrogen phosphate C11 H14 N3 O7 P KFCQHWOGBVCKHR-UKTHLTGXSA-N |  | ||
| PMP (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | D [auth A], F [auth B] | 4'-DEOXY-4'-AMINOPYRIDOXAL-5'-PHOSPHATE C8 H13 N2 O5 P ZMJGSOSNSPKHNH-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 61.968 | α = 90 |
| b = 90.028 | β = 90 |
| c = 100.29 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| Aimless | data scaling |
| PDB_EXTRACT | data extraction |
| CrysalisPro | data reduction |
| REFMAC | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Russian Science Foundation | Russian Federation | 19-14-00164 |