Synthesis and Biological Evaluation of C(13)/C(13')-Bis(desmethyl)disorazole Z.
Bold, C.P., Lucena-Agell, D., Oliva, M.A., Diaz, J.F., Altmann, K.H.(2023) Angew Chem Int Ed Engl 62: e202212190-e202212190
- PubMed: 36281761 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/anie.202212190
- Primary Citation Related Structures:
8AHM - PubMed Abstract:
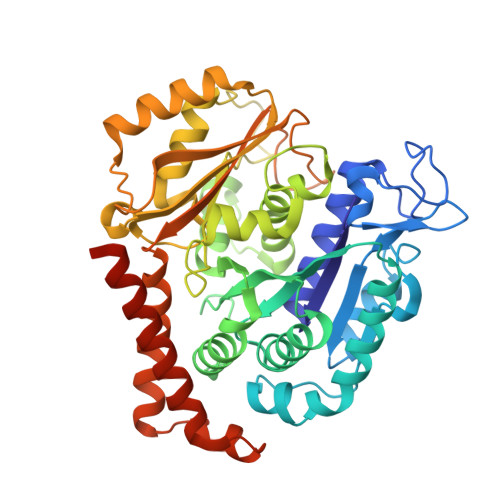
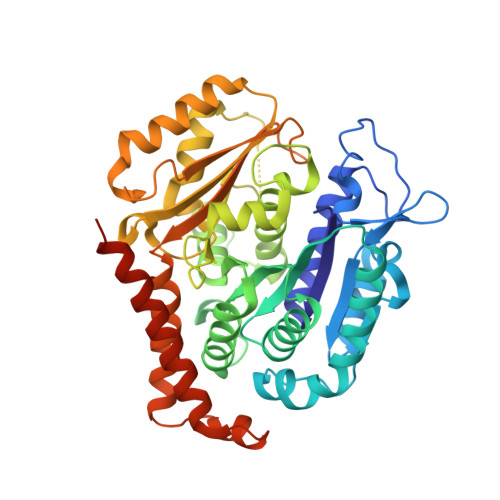
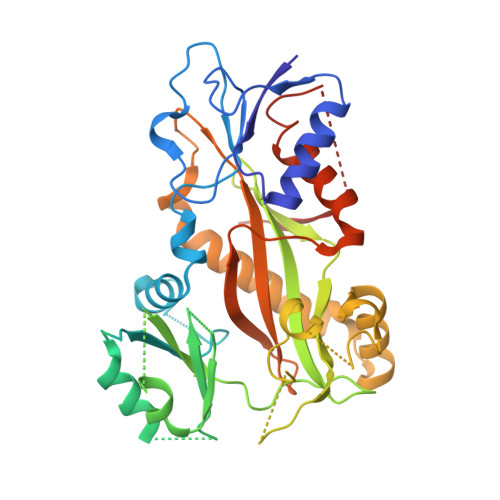
We describe the total synthesis of the macrodiolide C(13)/C(13')-bis(desmethyl)disorazole Z through double inter-/intramolecular Stille cross-coupling of a monomeric vinyl stannane/vinyl iodide precursor to form the macrocycle. The key step in the synthesis of this precursor was a stereoselective aldol reaction of a formal Evans acetate aldol product with crotonaldehyde. As demonstrated by X-ray crystallography, the binding mode of C(13)/C(13')-bis(desmethyl)disorazole Z to tubulin is virtually identical with that of the natural product disorazole Z. Likewise, C(13)/C(13')-bis(desmethyl)disorazole Z inhibits tubulin assembly with at least the same potency as disorazole Z and it appears to be a more potent cell growth inhibitor.
- Department of Chemistry and Applied Biosciences, Institute of Pharmaceutical Sciences, ETH Zürich, Vladimir-Prelog-Weg 4, 8093, Zürich, Switzerland.
Organizational Affiliation: