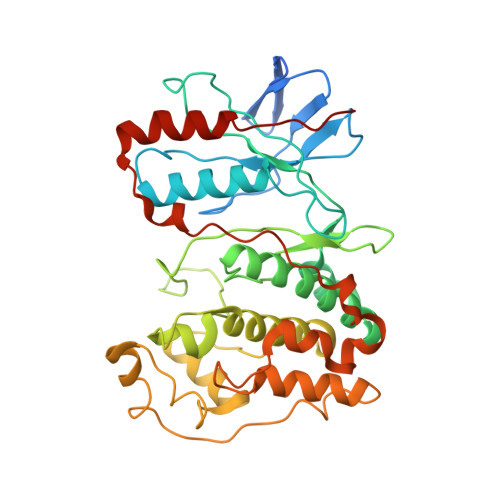
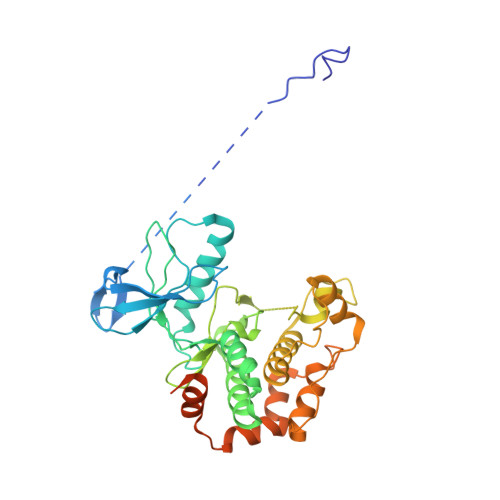
Architecture of the MKK6-p38 alpha complex defines the basis of MAPK specificity and activation.
Juyoux, P., Galdadas, I., Gobbo, D., von Velsen, J., Pelosse, M., Tully, M., Vadas, O., Gervasio, F.L., Pellegrini, E., Bowler, M.W.(2023) Science 381: 1217-1225
- PubMed: 37708276 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.add7859
- Primary Citation Related Structures:
8A8M - PubMed Abstract:
The mitogen-activated protein kinase (MAPK) p38α is a central component of signaling in inflammation and the immune response and is, therefore, an important drug target. Little is known about the molecular mechanism of its activation by double phosphorylation from MAPK kinases (MAP2Ks), because of the challenge of trapping a transient and dynamic heterokinase complex. We applied a multidisciplinary approach to generate a structural model of p38α in complex with its MAP2K, MKK6, and to understand the activation mechanism. Integrating cryo-electron microscopy with molecular dynamics simulations, hydrogen-deuterium exchange mass spectrometry, and experiments in cells, we demonstrate a dynamic, multistep phosphorylation mechanism, identify catalytically relevant interactions, and show that MAP2K-disordered amino termini determine pathway specificity. Our work captures a fundamental step of cell signaling: a kinase phosphorylating its downstream target kinase.
- European Molecular Biology Laboratory (EMBL), Grenoble, France.
Organizational Affiliation: