Extensive substrate recognition by the streptococcal antibody-degrading enzymes IdeS and EndoS.
Sudol, A.S.L., Butler, J., Ivory, D.P., Tews, I., Crispin, M.(2022) Nat Commun 13: 7801-7801
- PubMed: 36528711 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-35340-z
- Primary Citation Related Structures:
8A47, 8A48, 8A49 - PubMed Abstract:
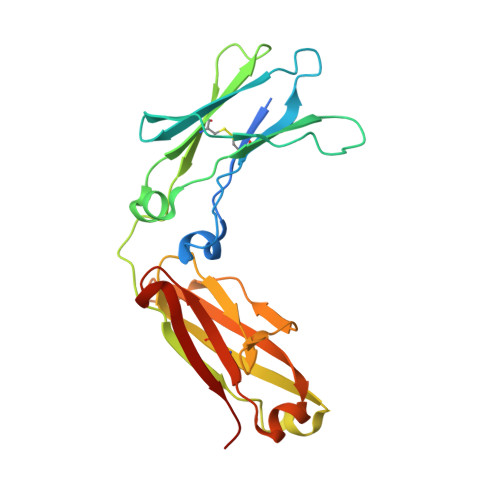
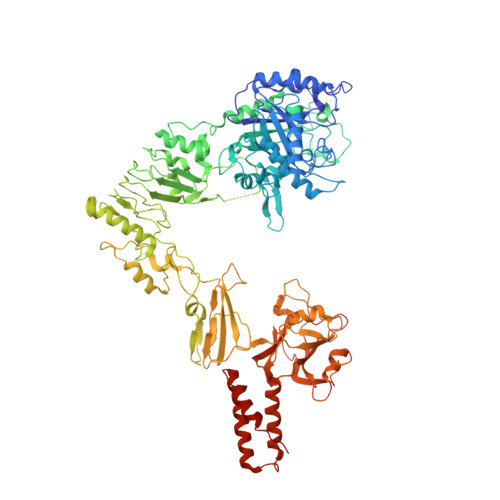
Enzymatic cleavage of IgG antibodies is a common strategy used by pathogenic bacteria to ablate immune effector function. The Streptococcus pyogenes bacterium secretes the protease IdeS and the glycosidase EndoS, which specifically catalyse cleavage and deglycosylation of human IgG, respectively. IdeS has received clinical approval for kidney transplantation in hypersensitised individuals, while EndoS has found application in engineering antibody glycosylation. We present crystal structures of both enzymes in complex with their IgG1 Fc substrate, which was achieved using Fc engineering to disfavour preferential Fc crystallisation. The IdeS protease displays extensive Fc recognition and encases the antibody hinge. Conversely, the glycan hydrolase domain in EndoS traps the Fc glycan in a "flipped-out" conformation, while additional recognition of the Fc peptide is driven by the so-called carbohydrate binding module. In this work, we reveal the molecular basis of antibody recognition by bacterial enzymes, providing a template for the development of next-generation enzymes.
- School of Biological Sciences, University of Southampton, Southampton, SO17 1BJ, UK.
Organizational Affiliation: