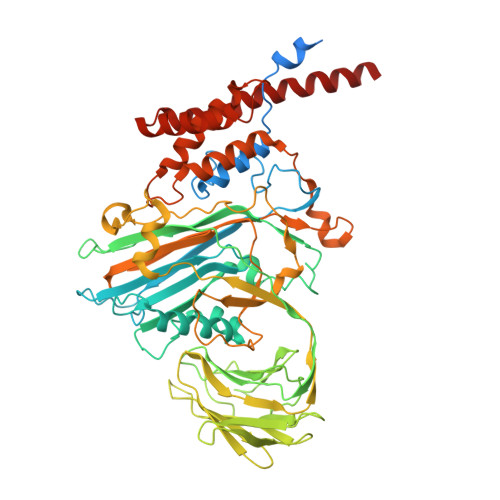
Unraveling the maturation pathway of a eukaryotic virus through cryo-EM.
Castells-Graells, R., Hesketh, E.L., Matsui, T., Johnson, J.E., Ranson, N.A., Lawson, D.M., Lomonossoff, G.P.(2026) Proc Natl Acad Sci U S A 123: e2420493123-e2420493123
- PubMed: 41739563 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2420493123
- Primary Citation Related Structures:
8A3C, 8A41, 8A6J, 8AAY, 8AC6, 8ACH - PubMed Abstract:
Virus maturation is a fundamental biological process involving large-scale structural reorganizations that drive functional activation and lead to infectivity. Understanding the steps from the initial procapsid assembly to mature virions is essential, both for comprehending viral life cycles and for developing antiviral therapies. However, capturing these steps has been challenging due to the transient and elusive nature of intermediate states. The nonenveloped, T = 4, ssRNA-containing, Nudaurelia capensis omega virus (NωV) is a highly accessible model system that exemplifies the maturation process of a eukaryotic virus. During maturation, the particle shrinks in outer diameter from 482 Å (pH 7.6) to 428 Å (pH 5.0). It is possible to mimic the maturation process in vitro by lowering the pH of a population of procapsids produced in heterologous systems. Indeed, by controlling the pH in vitro, it is possible to produce homogenous populations of intermediate NωV virus-like particles (VLPs) that occur too fleetingly to be observed in vivo. Here, we report structural models, based on cryoelectron microscopy (cryo-EM), of five intermediates in the NωV maturation process. The structures of the intermediate particles reveal unique, quaternary position-dependent trajectories and refolding of subunit N and C-terminal regions, including the formation of the autocatalytic cleavage site at N570. The detailed structures reported here, coupled with previously determined structures of the procapsids and mature particles, allow the maturation pathway to be described in detail for a eukaryotic virus.
- Department of Biochemistry and Metabolism, John Innes Centre, Norwich Research Park, Norwich NR4 7UH, United Kingdom.
Organizational Affiliation: