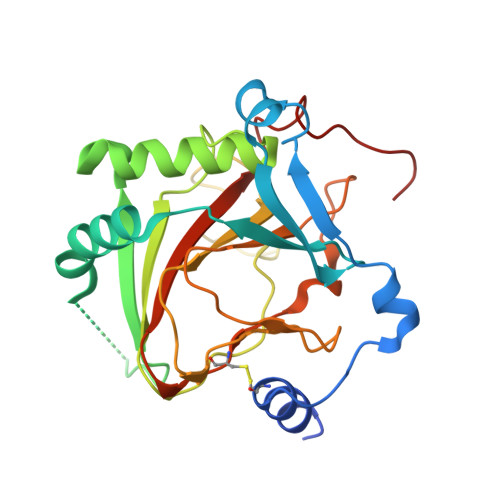
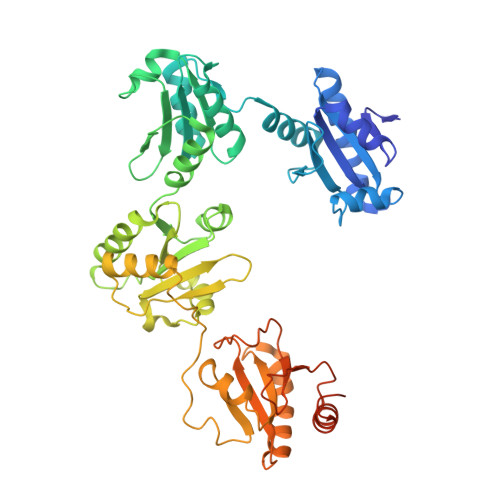
Crystal structure of the collagen prolyl 4-hydroxylase (C-P4H) catalytic domain complexed with PDI: Toward a model of the C-P4H alpha 2 beta 2 tetramer.
Murthy, A.V., Sulu, R., Lebedev, A., Salo, A.M., Korhonen, K., Venkatesan, R., Tu, H., Bergmann, U., Janis, J., Laitaoja, M., Ruddock, L.W., Myllyharju, J., Koski, M.K., Wierenga, R.K.(2022) J Biological Chem 298: 102614-102614
- PubMed: 36265586 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2022.102614
- Primary Citation Related Structures:
7ZSC - PubMed Abstract:
Collagen prolyl 4-hydroxylases (C-P4H) are α 2 β 2 tetramers, which catalyze the prolyl 4-hydroxylation of procollagen, allowing for the formation of the stable triple-helical collagen structure in the endoplasmic reticulum. The C-P4H α-subunit provides the N-terminal dimerization domain, the middle peptide-substrate-binding (PSB) domain, and the C-terminal catalytic (CAT) domain, whereas the β-subunit is identical to the enzyme protein disulfide isomerase (PDI). The structure of the N-terminal part of the α-subunit (N-terminal region and PSB domain) is known, but the structures of the PSB-CAT linker region and the CAT domain as well as its mode of assembly with the β/PDI subunit, are unknown. Here, we report the crystal structure of the CAT domain of human C-P4H-II complexed with the intact β/PDI subunit, at 3.8 Å resolution. The CAT domain interacts with the a, b', and a' domains of the β/PDI subunit, such that the CAT active site is facing bulk solvent. The structure also shows that the C-P4H-II CAT domain has a unique N-terminal extension, consisting of α-helices and a β-strand, which is the edge strand of its major antiparallel β-sheet. This extra region of the CAT domain interacts tightly with the β/PDI subunit, showing that the CAT-PDI interface includes an intersubunit disulfide bridge with the a' domain and tight hydrophobic interactions with the b' domain. Using this new information, the structure of the mature C-P4H-II α 2 β 2 tetramer is predicted. The model suggests that the CAT active-site properties are modulated by α-helices of the N-terminal dimerization domains of both subunits of the α 2 -dimer.
- Faculty of Biochemistry and Molecular Medicine, University of Oulu, Oulu, Finland; Biocenter Oulu, University of Oulu, Oulu, Finland.
Organizational Affiliation: