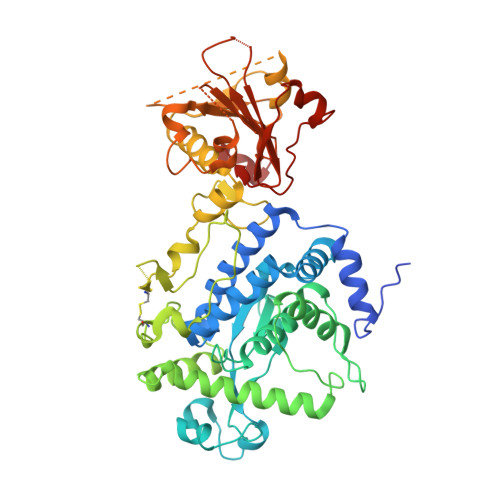
Unconventional structure and mechanisms for membrane interaction and translocation of the NF-kappa B-targeting toxin AIP56.
Lisboa, J., Pereira, C., Pinto, R.D., Rodrigues, I.S., Pereira, L.M.G., Pinheiro, B., Oliveira, P., Pereira, P.J.B., Azevedo, J.E., Durand, D., Benz, R., do Vale, A., Dos Santos, N.M.S.(2023) Nat Commun 14: 7431-7431
- PubMed: 37973928 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-43054-z
- Primary Citation Related Structures:
7ZPF - PubMed Abstract:
Bacterial AB toxins are secreted key virulence factors that are internalized by target cells through receptor-mediated endocytosis, translocating their enzymatic domain to the cytosol from endosomes (short-trip) or the endoplasmic reticulum (long-trip). To accomplish this, bacterial AB toxins evolved a multidomain structure organized into either a single polypeptide chain or non-covalently associated polypeptide chains. The prototypical short-trip single-chain toxin is characterized by a receptor-binding domain that confers cellular specificity and a translocation domain responsible for pore formation whereby the catalytic domain translocates to the cytosol in an endosomal acidification-dependent way. In this work, the determination of the three-dimensional structure of AIP56 shows that, instead of a two-domain organization suggested by previous studies, AIP56 has three-domains: a non-LEE encoded effector C (NleC)-like catalytic domain associated with a small middle domain that contains the linker-peptide, followed by the receptor-binding domain. In contrast to prototypical single-chain AB toxins, AIP56 does not comprise a typical structurally complex translocation domain; instead, the elements involved in translocation are scattered across its domains. Thus, the catalytic domain contains a helical hairpin that serves as a molecular switch for triggering the conformational changes necessary for membrane insertion only upon endosomal acidification, whereas the middle and receptor-binding domains are required for pore formation.
- Fish Immunology and Vaccinology Group, IBMC-Instituto de Biologia Molecular e Celular, Universidade do Porto, 4200-135, Porto, Portugal. johnny.lisboa@i3s.up.pt.
Organizational Affiliation: