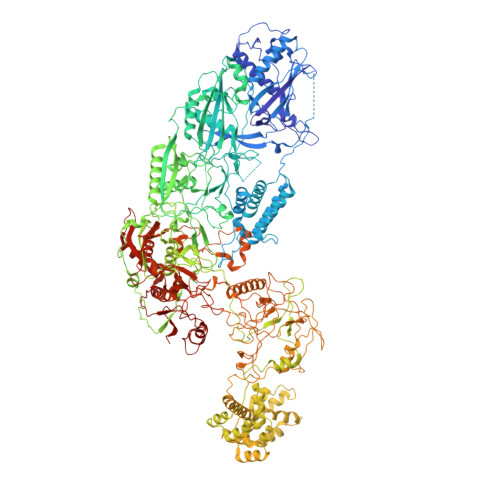
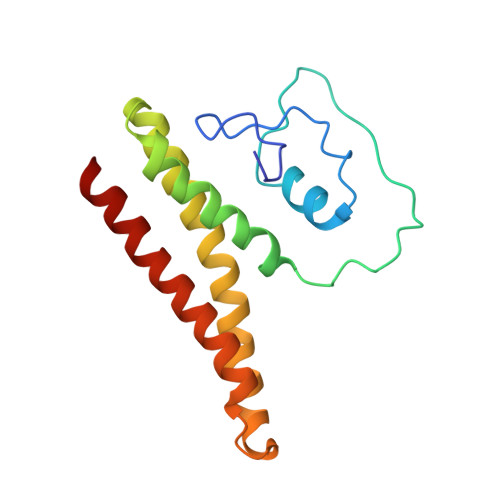
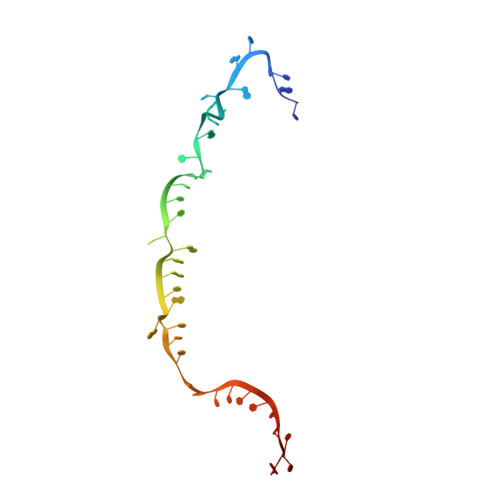
Structural insights into the regulation of Cas7-11 by TPR-CHAT.
Ekundayo, B., Torre, D., Beckert, B., Nazarov, S., Myasnikov, A., Stahlberg, H., Ni, D.(2023) Nat Struct Mol Biol 30: 135-139
- PubMed: 36471056 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-022-00894-5
- Primary Citation Related Structures:
7ZOL, 7ZOQ - PubMed Abstract:
The CRISPR-guided caspase (Craspase) complex is an assembly of the target-specific RNA nuclease known as Cas7-11 bound to CRISPR RNA (crRNA) and an ancillary protein known as TPR-CHAT (tetratricopeptide repeats (TPR) fused with a CHAT domain). The Craspase complex holds promise as a tool for gene therapy and biomedical research, but its regulation is poorly understood. TPR-CHAT regulates Cas7-11 nuclease activity via an unknown mechanism. In the present study, we use cryoelectron microscopy to determine structures of the Desulfonema magnum (Dm) Craspase complex to gain mechanistic insights into its regulation. We show that DmTPR-CHAT stabilizes crRNA-bound DmCas7-11 in a closed conformation via a network of interactions mediated by the DmTPR-CHAT N-terminal domain, the DmCas7-11 insertion finger and Cas11-like domain, resulting in reduced target RNA accessibility and cleavage.
- Laboratory of Biological Electron Microscopy, Institute of Physics, School of Basic Sciences, EPFL, and Department of Fundamental Microbiology, Faculty of Biology and Medicine, University of Lausanne, Lausanne, Switzerland.
Organizational Affiliation: