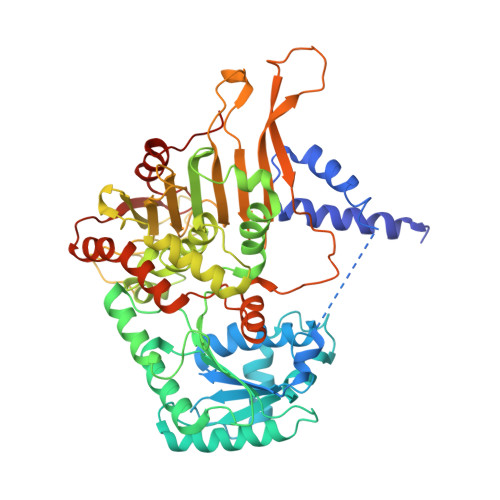
Crystal structure of Leishmania donovani glucose 6-phosphate dehydrogenase reveals a unique N-terminal domain.
Berneburg, I., Rahlfs, S., Becker, K., Fritz-Wolf, K.(2022) Commun Biol 5: 1353-1353
- PubMed: 36494598 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-022-04307-7
- Primary Citation Related Structures:
7ZHT, 7ZHU, 7ZHV, 7ZHW, 7ZHX, 7ZHY, 7ZHZ - PubMed Abstract:
Since unicellular parasites highly depend on NADPH as a source for reducing equivalents, the pentose phosphate pathway, especially the first and rate-limiting NADPH-producing enzyme glucose 6-phosphate dehydrogenase (G6PD), is considered an excellent antitrypanosomatid drug target. Here we present the crystal structure of Leishmania donovani G6PD (LdG6PD) elucidating the unique N-terminal domain of Kinetoplastida G6PDs. Our investigations on the function of the N-domain suggest its involvement in the formation of a tetramer that is completely different from related Trypanosoma G6PDs. Structural and functional investigations further provide interesting insights into the binding mode of LdG6PD, following an ordered mechanism, which is confirmed by a G6P-induced domain shift and rotation of the helical N-domain. Taken together, these insights into LdG6PD contribute to the understanding of G6PDs' molecular mechanisms and provide an excellent basis for further drug discovery approaches.
- Biochemistry and Molecular Biology, Interdisciplinary Research Center, Justus Liebig University, Giessen, Germany.
Organizational Affiliation: