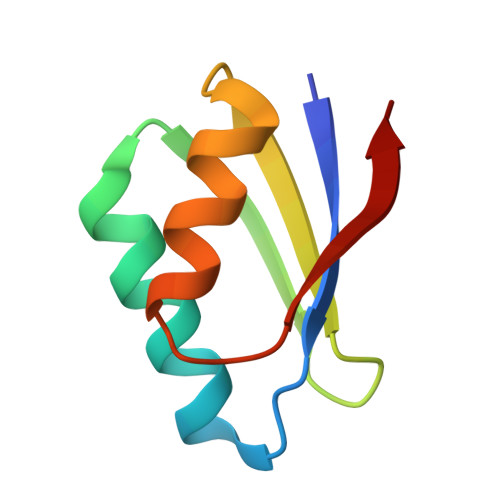
Crystal Structure of the Human Copper Chaperone ATOX1 Bound to Zinc Ion.
Mangini, V., Belviso, B.D., Nardella, M.I., Natile, G., Arnesano, F., Caliandro, R.(2022) Biomolecules 12
- PubMed: 36291703 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/biom12101494
- Primary Citation Related Structures:
7ZC3 - PubMed Abstract:
The bioavailability of copper (Cu) in human cells may depend on a complex interplay with zinc (Zn) ions. We investigated the ability of the Zn ion to target the human Cu-chaperone Atox1, a small cytosolic protein capable of anchoring Cu(I), by a conserved surface-exposed Cys-X-X-Cys (CXXC) motif, and deliver it to Cu-transporting ATPases in the trans-Golgi network. The crystal structure of Atox1 loaded with Zn displays the metal ion bridging the CXXC motifs of two Atox1 molecules in a homodimer. The identity and location of the Zn ion were confirmed through the anomalous scattering of the metal by collecting X-ray diffraction data near the Zn K-edge. Furthermore, soaking experiments of the Zn-loaded Atox1 crystals with a strong chelating agent, such as EDTA, caused only limited removal of the metal ion from the tetrahedral coordination cage, suggesting a potential role of Atox1 in Zn metabolism and, more generally, that Cu and Zn transport mechanisms could be interlocked in human cells.
- Institute of Crystallography, CNR, Via G. Amendola 122/o, 70126 Bari, Italy.
Organizational Affiliation: