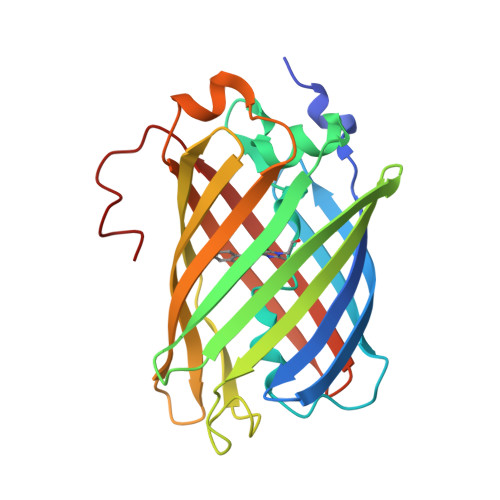
Cyan fluorescent proteins derived from mNeonGreen.
Zarowny, L., Clavel, D., Johannson, R., Duarte, K., Depernet, H., Dupuy, J., Baker, H., Brown, A., Royant, A., Campbell, R.E.(2022) Protein Eng Des Sel 35
- PubMed: 35417013 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/protein/gzac004
- Primary Citation Related Structures:
7Z7O, 7Z7P, 7Z7Q - PubMed Abstract:
mNeonGreen, an engineered green fluorescent protein (GFP) derived from lancelet, is one of the most brightly fluorescent homologs of Aequorea victoria jellyfish GFP (avGFP) yet reported. In this work, we investigated whether this bright fluorescence might be retained in homologs of mNeonGreen with modified chromophore structures and altered fluorescent hues. We found mNeonGreen to be generally less tolerant than avGFP to chromophore modification by substitution of the key chromophore-forming tyrosine residue with other aromatic amino acids. However, we were ultimately successful in creating a variant, designated as NeonCyan1, with a tryptophan-derived cyan fluorescent protein (CFP)-type chromophore, and two additional mutants with distinct spectral hues. Structural, computational, and photophysical characterization of NeonCyan1 and its variants provided insight into the factors that control the fluorescence emission color. Though not recommended as replacements for contemporary CFP variants, we demonstrate that NeonCyan1 variants are potentially suitable for live cell imaging applications.
- Department of Chemistry, University of Alberta, Edmonton T6G 2G2, Canada.
Organizational Affiliation: