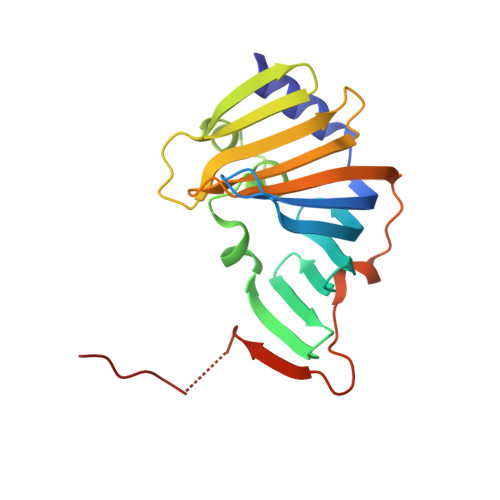
Structural basis of lipoprotein recognition by the bacterial Lol trafficking chaperone LolA.
Kaplan, E., Greene, N.P., Jepson, A.E., Koronakis, V.(2022) Proc Natl Acad Sci U S A 119: e2208662119-e2208662119
- PubMed: 36037338 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2208662119
- Primary Citation Related Structures:
7Z6W, 7Z6X - PubMed Abstract:
In gram-negative bacteria, lipoproteins are vital structural components of the outer membrane (OM) and crucial elements of machineries central to the physiology of the cell envelope. A dedicated apparatus, the Lol system, is required for the correct localization of OM lipoproteins and is essential for viability. The periplasmic chaperone LolA is central to this trafficking pathway, accepting triacylated lipoproteins from the inner membrane transporter LolCDE, before carrying them across the periplasm to the OM receptor LolB. Here, we report a crystal structure of liganded LolA, generated in vivo, revealing the molecular details of lipoprotein association. The structure highlights how LolA, initially primed to receive lipoprotein by interaction with LolC, further opens to accommodate the three ligand acyl chains in a precise conformation within its cavity. LolA forms extensive interactions with the acyl chains but not with any residue of the cargo, explaining the chaperone's ability to transport structurally diverse lipoproteins. Structural characterization of a ligandedLolA variant incapable of lipoprotein release reveals aberrant association, demonstrating the importance of the LolCDE-coordinated, sequential opening of LolA for inserting lipoprotein in a manner productive for subsequent trafficking. Comparison with existing structures of LolA in complex with LolC or LolCDE reveals substantial overlap of the lipoprotein and LolC binding sites within the LolA cavity, demonstrating that insertion of lipoprotein acyl chains physically disengages the chaperone protein from the transporter by perturbing interaction with LolC. Taken together, our data provide a key step toward a complete understanding of a fundamentally important trafficking pathway.
- Department of Pathology, University of Cambridge, Cambridge CB2 1QP, United Kingdom.
Organizational Affiliation: