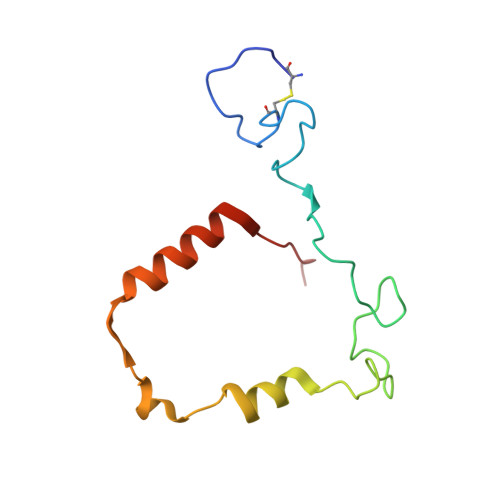
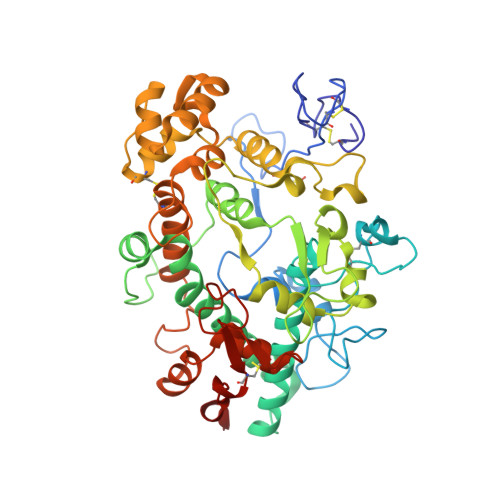
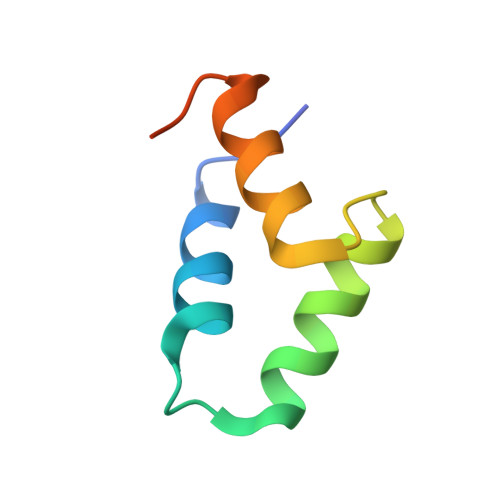
The staphylococcal inhibitory protein SPIN binds to human myeloperoxidase with picomolar affinity but only dampens halide oxidation.
Leitgeb, U., Furtmuller, P.G., Hofbauer, S., Brito, J.A., Obinger, C., Pfanzagl, V.(2022) J Biological Chem 298: 102514-102514
- PubMed: 36150500 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2022.102514
- Primary Citation Related Structures:
7QZR, 7Z53 - PubMed Abstract:
The heme enzyme myeloperoxidase (MPO) is one of the key players in the neutrophil-mediated killing of invading pathogens as part of the innate immune system. MPO generates antimicrobial oxidants, which indiscriminately and effectively kill phagocytosed pathogens. Staphylococcus aureus, however, is able to escape this fate, in part by secreting a small protein called SPIN (Staphylococcal Peroxidase Inhibitor), which specifically targets and inhibits MPO in a structurally complex manner. Here, we present the first crystal structures of the complex of SPIN-aureus and a truncated version (SPIN-truncated) with mature dimeric leukocyte MPO. We unravel the contributions of the two domains to the kinetics and thermodynamics of SPIN-aureus binding to MPO by using a broad array of complementary biochemical and biophysical methods. The C-terminal "recognition" domain is shown to mediate specific binding to MPO, while interaction of the N-terminal "inhibitory" domain is guided mainly by hydrophobic effects and thus is less sequence dependent. We found that inhibition of MPO is achieved by reducing substrate migration, but SPIN-aureus cannot completely block MPO activity. Its' effectiveness is inversely related to substrate size, with no discernible dependence on other factors. Thus, SPIN-aureus is an extremely high-affinity inhibitor and highly efficient for substrates larger than halogens. As aberrant MPO activity is implicated in a number of chronic inflammatory diseases, SPIN-aureus is the first promising protein inhibitor for specific inhibition of human MPO.
- University of Natural Resources and Life Sciences, Vienna, Department of Chemistry, Institute of Biochemistry, Vienna, Austria.
Organizational Affiliation: