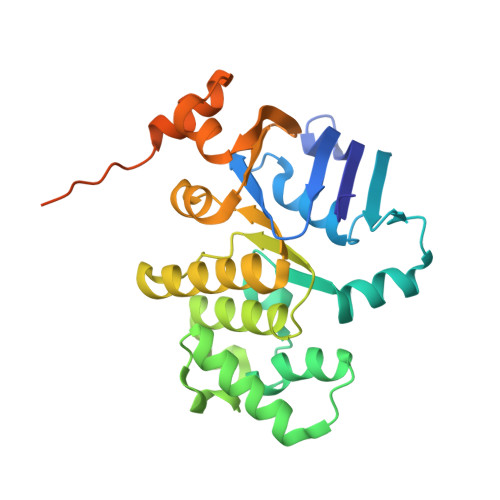
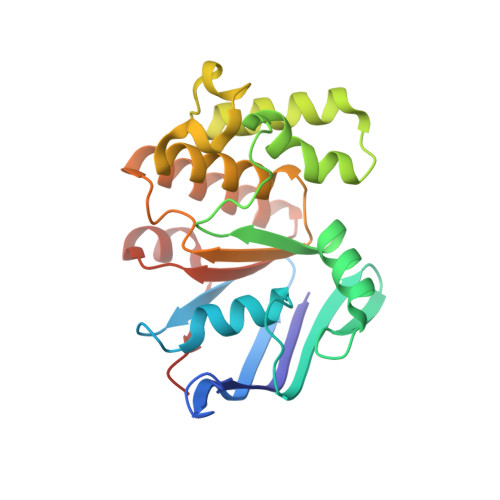
Structural remodelling of the carbon-phosphorus lyase machinery by a dual ABC ATPase.
Amstrup, S.K., Ong, S.C., Sofos, N., Karlsen, J.L., Skjerning, R.B., Boesen, T., Enghild, J.J., Hove-Jensen, B., Brodersen, D.E.(2023) Nat Commun 14: 1001-1001
- PubMed: 36813778 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-36604-y
- Primary Citation Related Structures:
7Z15, 7Z16, 7Z17, 7Z18, 7Z19 - PubMed Abstract:
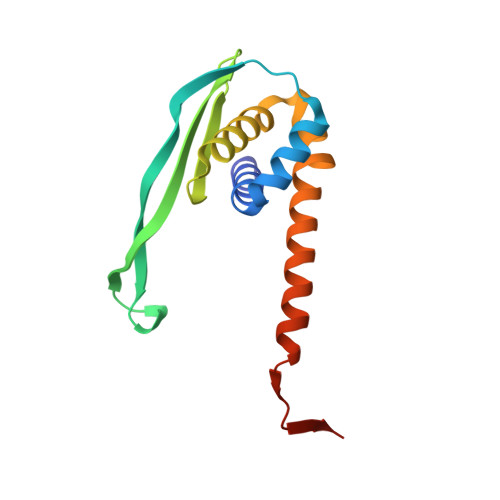
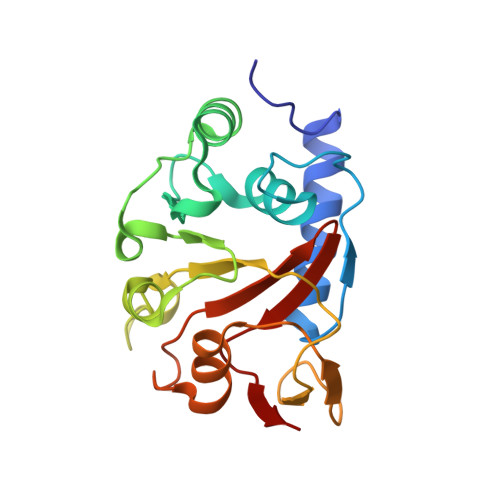
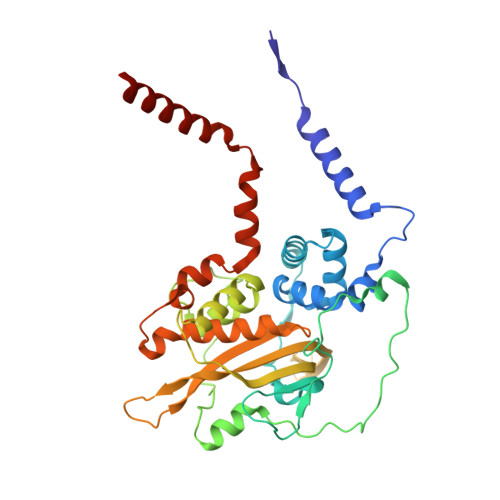
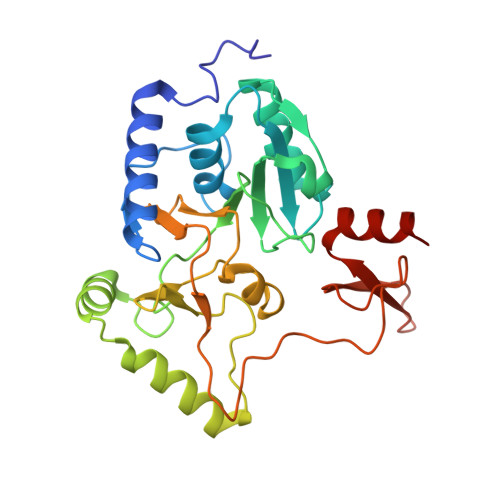
In Escherichia coli, the 14-cistron phn operon encoding carbon-phosphorus lyase allows for utilisation of phosphorus from a wide range of stable phosphonate compounds containing a C-P bond. As part of a complex, multi-step pathway, the PhnJ subunit was shown to cleave the C-P bond via a radical mechanism, however, the details of the reaction could not immediately be reconciled with the crystal structure of a 220 kDa PhnGHIJ C-P lyase core complex, leaving a significant gap in our understanding of phosphonate breakdown in bacteria. Here, we show using single-particle cryogenic electron microscopy that PhnJ mediates binding of a double dimer of the ATP-binding cassette proteins, PhnK and PhnL, to the core complex. ATP hydrolysis induces drastic structural remodelling leading to opening of the core complex and reconfiguration of a metal-binding and putative active site located at the interface between the PhnI and PhnJ subunits.
- Department of Molecular Biology and Genetics, Aarhus University, Universitetsbyen 81, DK-8000, Aarhus C, Denmark.
Organizational Affiliation: