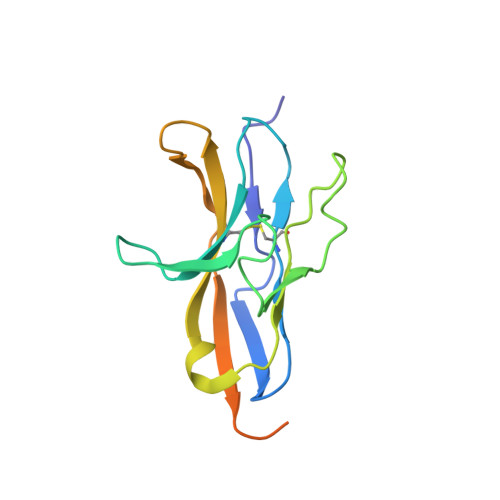
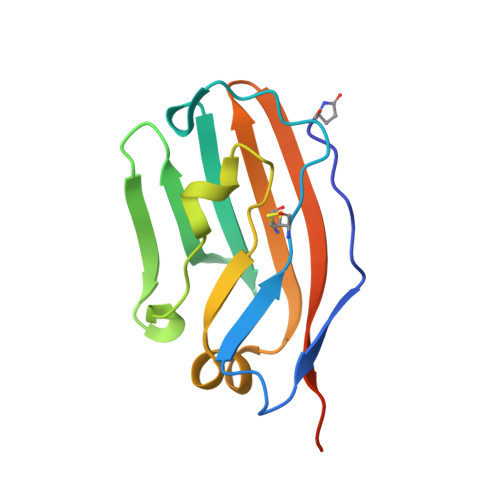
Crystal Structure of Human CD47 in Complex with Engineered SIRP alpha.D1(N80A).
Yu, J., Li, S., Chen, D., Liu, D., Guo, H., Yang, C., Zhang, W., Zhang, L., Zhao, G., Tu, X., Peng, L., Liu, S., Bai, X., Song, Y., Jiang, Z., Zhang, R., Tian, W.(2022) Molecules 27
- PubMed: 36080360
- DOI: https://doi.org/10.3390/molecules27175574
- Primary Citation Related Structures:
7YGG - PubMed Abstract:
Background: Targeting the CD47/SIRPα signaling pathway represents a novel approach to enhance anti-tumor immunity. However, the crystal structure of the CD47/SIRPα has not been fully studied. This study aims to analyze the structure interface of the complex of CD47 and IMM01, a novel recombinant SIRPα-Fc fusion protein. Methods: IMM01-Fab/CD47 complex was crystalized, and diffraction images were collected. The complex structure was determined by molecular replacement using the program PHASER with the CD47-SIRPαv2 structure (PDB code 2JJT) as a search model. The model was manually built using the COOT program and refined using TLS parameters in REFMAC from the CCP4 program suite. Results: Crystallization and structure determination analysis of the interface of IMM01/CD47 structure demonstrated CD47 surface buried by IMM01. Comparison with the literature structure (PDB ID 2JJT) showed that the interactions of IMM01/CD47 structure are the same. All the hydrogen bonds that appear in the literature structure are also present in the IMM01/CD47 structure. These common hydrogen bonds are stable under different crystal packing styles, suggesting that these hydrogen bonds are important for protein binding. In the structure of human CD47 in complex with human SIRPα, except SER66, the amino acids that form hydrogen bonds are all conserved. Furthermore, comparing with the structure of PDB ID 2JJT, the salt bridge interaction from IMM01/CD47 structure are very similar, except the salt bridge bond between LYS53 in IMM01 and GLU106 in CD47, which only occurs between the B and D chains. However, as the side chain conformation of LYS53 in chain A is slightly different, the salt bridge bond is absent between the A and C chains. At this site between chain A and chain C, there are a salt bridge bond between LYS53 (A) and GLU104 (C) and a salt bridge bond between HIS56 (A) and GLU106 (C) instead. According to the sequence alignment results of SIRPα, SIRPβ and SIRPγ in the literature of PDB ID 2JJT, except ASP100, the amino acids that form common salt bridge bonds are all conserved. Conclusion: Our data demonstrated crystal structure of the IMM01/CD47 complex and provides a structural basis for the structural binding interface and future clinical applications.
- Department of Hematology, The First Affiliated Hospital of Zhengzhou University, Zhengzhou 450052, China.
Organizational Affiliation: