Highly Potent and Oral Macrocyclic Peptides as a HIV-1 Protease Inhibitor: mRNA Display-Derived Hit-to-Lead Optimization.
Kusumoto, Y., Hayashi, K., Sato, S., Yamada, T., Kozono, I., Nakata, Z., Asada, N., Mitsuki, S., Watanabe, A., Wakasa-Morimoto, C., Uemura, K., Arita, S., Miki, S., Mizutare, T., Mikamiyama, H.(2022) ACS Med Chem Lett 13: 1634-1641
- PubMed: 36262395 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsmedchemlett.2c00310
- Primary Citation Related Structures:
7YF6 - PubMed Abstract:
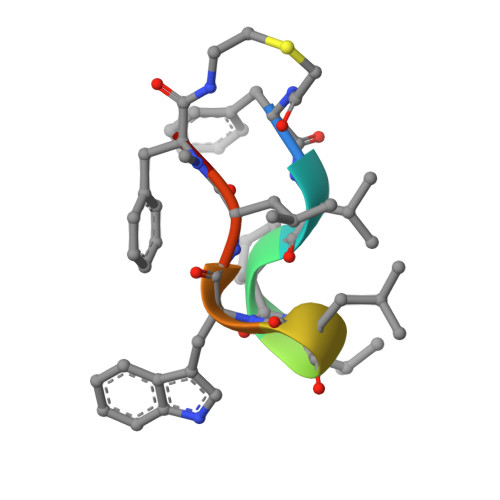
Human immunodeficiency virus type-1 (HIV-1) protease is essential for viral propagation, and its inhibitors are key anti-HIV-1 drug candidates. In this study, we discovered a novel HIV-1 protease inhibitor (compound 16 ) with potent antiviral activity and oral bioavailability using a structure-based drug design approach via X-ray crystal structure analysis and improved metabolic stability, starting from hit macrocyclic peptides identified by mRNA display against HIV-1 protease. We found that the improvement of the proteolytic stability of macrocyclic peptides by introducing a methyl group to the α-position of amino acid is crucial to exhibit strong antiviral activity. In addition, macrocyclic peptides, which have moderate metabolic stability and solubility in solutions containing taurocholic acid, exhibited desirable plasma total clearance and oral bioavailability. These approaches may contribute to the successful discovery and development of orally bioavailable peptide drugs.
- Shionogi Pharmaceutical Research Center, Shionogi & Co., Ltd. 1-1, Futaba-cho 3-chome, Toyonaka, Osaka 561-0825, Japan.
Organizational Affiliation: