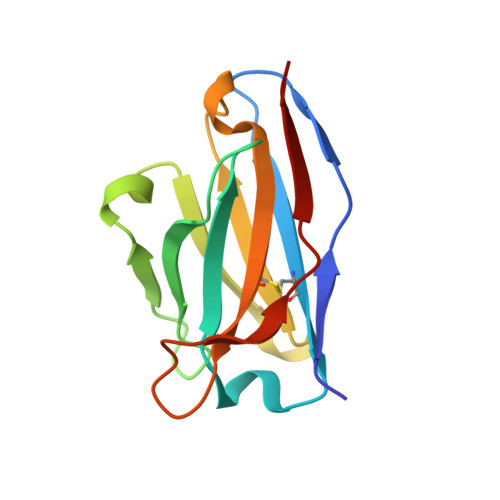
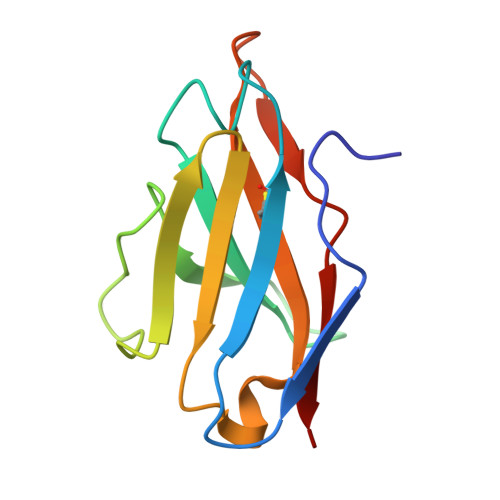
A broadly neutralizing antibody recognizes a unique epitope with a signature motif common across coronaviruses
Yan, L., Wang, F., Hill, M., Brun, J., Liang, Z., Shi, X., Zhang, L., He, X., Li, Y., Huang, Q., Dong, X., Liu, H., Zhang, Y., Liu, L., Dwek, R.A., Zitzmann, N., Liang, A., Yang, G.(2025) Nat Commun 16: 7580