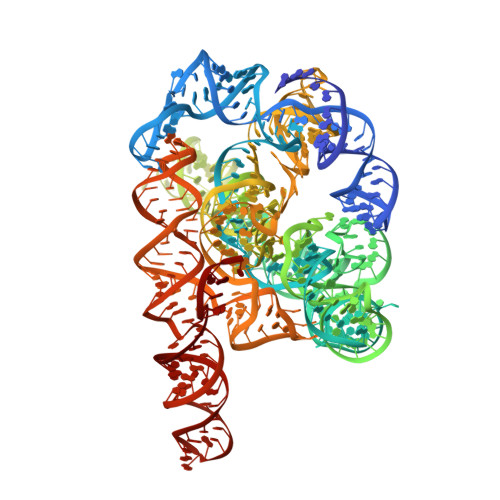
Snapshots of the first-step self-splicing of Tetrahymena ribozyme revealed by cryo-EM.
Zhang, X., Li, S., Pintilie, G., Palo, M.Z., Zhang, K.(2023) Nucleic Acids Res 51: 1317-1325
- PubMed: 36660826 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkac1268
- Primary Citation Related Structures:
7YC8, 7YCG, 7YCH, 7YCI - PubMed Abstract:
Tetrahymena ribozyme is a group I intron, whose self-splicing is the result of two sequential ester-transfer reactions. To understand how it facilitates catalysis in the first self-splicing reaction, we used cryogenic electron microscopy (cryo-EM) to resolve the structures of L-16 Tetrahymena ribozyme complexed with a 11-nucleotide 5'-splice site analog substrate. Four conformations were achieved to 4.14, 3.18, 3.09 and 2.98 Å resolutions, respectively, corresponding to different splicing intermediates during the first enzymatic reaction. Comparison of these structures reveals structural alterations, including large conformational changes in IGS/IGSext (P1-P1ext duplex) and J5/4, as well as subtle local rearrangements in the G-binding site. These structural changes are required for the enzymatic activity of the Tetrahymena ribozyme. Our study demonstrates the ability of cryo-EM to capture dynamic RNA structural changes, ushering in a new era in the analysis of RNA structure-function by cryo-EM.
- Department of Urology, The First Affiliated Hospital of USTC, MOE Key Laboratory for Cellular Dynamics, Division of Life Sciences and Medicine, University of Science and Technology of China, Hefei 230001, China.
Organizational Affiliation: