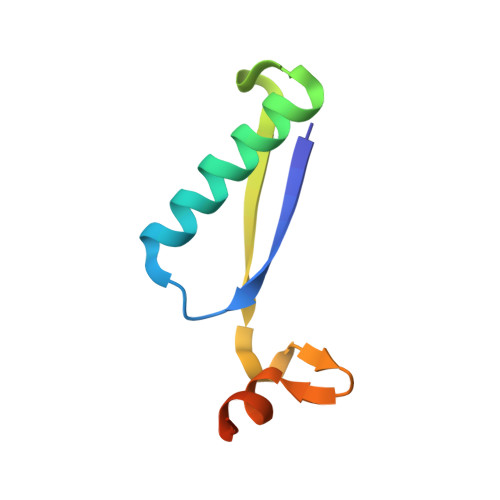
Structural insights into the substrate specificity of 5-chloro-2-hydroxymuconate tautomerase CnbG.
Ma, H.L., Ding, M., Guo, L., Li, D.F.(2022) Biochem Biophys Res Commun 620: 42-48
- PubMed: 35777133 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2022.06.058
- Primary Citation Related Structures:
7XUY - PubMed Abstract:
The catechol meta-cleavage pathway is widely involved in the degradation of aromatic compounds, including those halogenated aromatic hydrocarbons and their derivatives. CnbG is a kind of 4-oxalocrotonate tautomerase (4-OT) located in the catechol meta-cleavage pathway, catalyzes the ketonization of cis,cis-5-chloro-2-hydroxymuconate and cis,cis-2-hydroxymuconate to yield 5-chloro-2-oxo-3-hexene-1,6-dioate and 2-oxo-3-hexene-1,6-dioate, and contributes to the degradation of 4-chloronitrobenzene and chlorobenzene in Comamonas testosteroni CNB-1. Yet, the reason why CnbG and those 4-OTs could recognize various substrates is not well explained. Here, we determined the crystal structure of CnbG at resolution of 2.0 Å and identified that the potential substrate pocket involved in four conserved residues, residues Pro1, Arg11, Arg39 and Trp50, but not five conserved residues as those reported in other 4-OTs. We also found the four conserved residues assemble different sequence patterns in different 4-OTs, indicating their potential roles in catalysis and substrate binding. Via molecular docking, we found the 5-chloro group was clamped by two residues and extended to the solvent, indicating a substrate binding mode that could bear the substitution of different groups in the 5-position. Our work extends the knowledge of the substrate specificity of enzymes in the catechol meta-cleavage pathway.
- State Key Laboratory of Microbial Resources, Institute of Microbiology, Chinese Academy of Sciences, Beijing, 100101, PR China; University of Chinese Academy of Sciences, Beijing, 100049, PR China.
Organizational Affiliation: