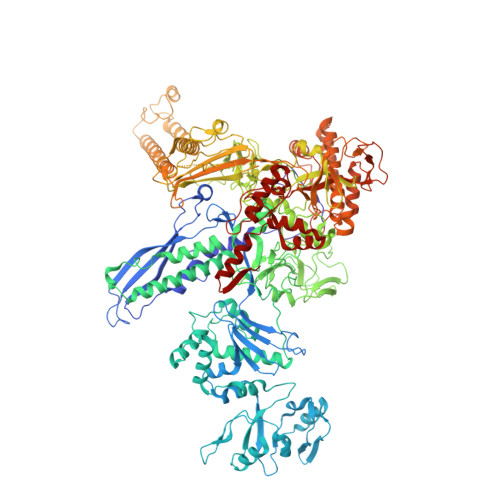
Structural basis of transcriptional regulation by a nascent RNA element, HK022 putRNA.
Hwang, S., Olinares, P.D.B., Lee, J., Kim, J., Chait, B.T., King, R.A., Kang, J.Y.(2022) Nat Commun 13: 4668-4668
- PubMed: 35970830 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-32315-y
- Primary Citation Related Structures:
7XUE, 7XUG, 7XUI - PubMed Abstract:
Transcription, in which RNA polymerases (RNAPs) produce RNA from DNA, is the first step of gene expression. As such, it is highly regulated either by trans-elements like protein factors and/or by cis-elements like specific sequences on the DNA. Lambdoid phage HK022 contains a cis-element, put, which suppresses pausing and termination during transcription of the early phage genes. The putRNA transcript solely performs the anti-pausing/termination activities by interacting directly with the E.coli RNAP elongation complex (EC) by an unknown structural mechanism. In this study, we reconstituted putRNA-associated ECs and determined the structures using cryo-electron microscopy. The determined structures of putRNA-associated EC, putRNA-absent EC, and σ 70 -bound EC suggest that the putRNA interaction with the EC counteracts swiveling, a conformational change previously identified to promote pausing and σ 70 might modulate putRNA folding via σ 70 -dependent pausing during elongation.
- Department of Chemistry, Korea Advanced Institute of Science and Technology (KAIST), Daejeon, Republic of Korea.
Organizational Affiliation: