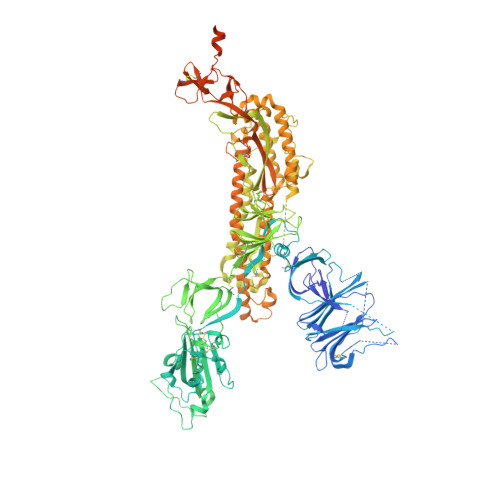
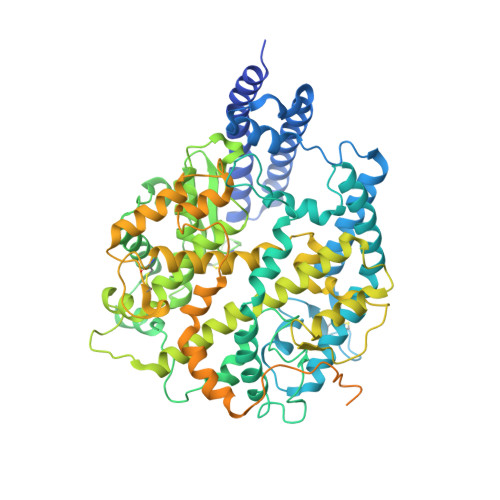
Structural and functional analysis of an inter-Spike bivalent neutralizing antibody against SARS-CoV-2 variants.
Li, Y., Fan, Q., Zhou, B., Shen, Y., Zhang, Y., Cheng, L., Qi, F., Song, S., Guo, Y., Yan, R., Ju, B., Zhang, Z.(2022) iScience 25: 104431-104431
- PubMed: 35607524 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.isci.2022.104431
- Primary Citation Related Structures:
7XIC, 7XID - PubMed Abstract:
The different variants of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) have attracted most public concern because they caused "wave and wave" COVID-19 pandemic. The initial step of viral infection is mediated by the SARS-CoV-2 Spike (S) protein, which mediates the receptor recognition and membrane fusion between virus and host cells. Neutralizing antibodies (nAbs) targeting the S protein of SARS-CoV-2 have become promising candidates for clinical intervention strategy, while multiple studies have shown that different variants have enhanced infectivity and antibody resistance. Here, we explore the structure and function of STS165, a broadly inter-Spike bivalent nAb against SARS-CoV-2 variants and even SARS-CoV, contributing to further understanding of the working mechanism of nAbs.
- Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Zhejiang Province, Hangzhou 310024, China.
Organizational Affiliation: