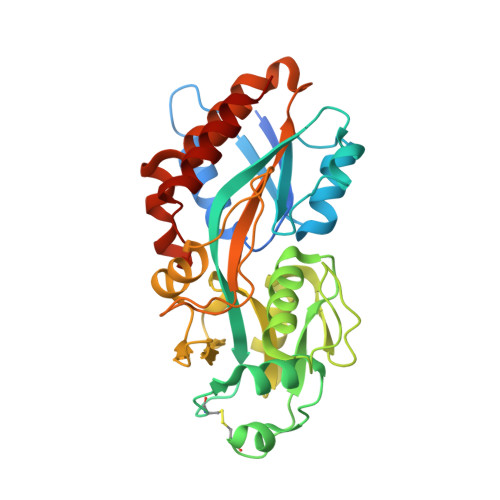
Biochemical and structural characterization of the cyanophage-encoded phosphate-binding protein: implications for enhanced phosphate uptake of infected cyanobacteria
Zhao, F., Lin, X., Cai, K., Jiang, Y., Ni, T., Chen, Y., Feng, J., Dang, S., Zhou, C.Z., Zeng, Q.(2022) Environ Microbiol 24: 3037-3050