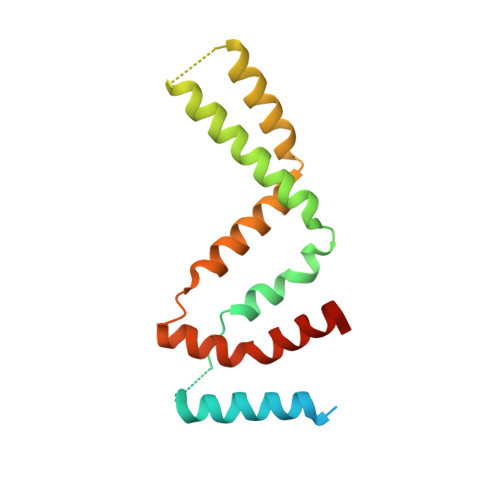
Novel Dimeric Architecture of an IFN-gamma-Related Cytokine Provides Insights into Subfunctionalization of Type II IFNs in Teleost Fish.
Zhu, X., Wang, J., Jia, Z., Feng, J., Wang, B., Wang, Z., Liu, Q., Wu, K., Huang, W., Zhao, X., Dang, H., Zou, J.(2022) J Immunol 209: 2203-2214
- PubMed: 36426983 Search on PubMed
- DOI: https://doi.org/10.4049/jimmunol.2200334
- Primary Citation Related Structures:
7X45 - PubMed Abstract:
Gene duplication leads to subfunctionalization of paralogs. In mammals, IFN-γ is the sole member of the type II IFN family and binds to a receptor complex consisting of IFN-γR1 and IFN-γR2. In teleost fish, IFN-γ and its receptors have been duplicated due to the teleost-specific whole-genome duplication event. In this study, the functions of an IFN-γ-related (IFN-γrel) cytokine were found to be partially retained relative to IFN-γ in grass carp (Ctenopharyngodon idella [CiIFN-γrel]). CiIFN-γrel upregulated the expression of proinflammatory genes but had lost the ability to activate genes involved in Th1 response. The results suggest that CiIFN-γrel could have been subfunctionalized from CiIFN-γ. Moreover, CiIFN-γrel induced STAT1 phosphorylation via interaction with duplicated homologs of IFN-γR1 (cytokine receptor family B [CRFB] 17 and CRFB13). Strikingly, CiIFN-γrel did not bind to the IFN-γR2 homolog (CRFB6). To gain insight into the subfunctionalization, the crystal structure of CiIFN-γrel was solved at 2.26 Å, revealing that it forms a homodimer that is connected by two pairs of disulfide bonds. Due to the spatial positions of helix A, loop AB, and helix B, CiIFN-γrel displays a unique topology that requires elements from two identical monomers to form a unit that is similar to IFN-γ. Further, mutagenesis analyses identified key residues interacting with CiIFN-γrel receptors and those required for the biological functions. Our study can help understand the subfunctionalization of duplicated IFN-γ paralogs in fish.
- Key Laboratory of Exploration and Utilization of Aquatic Genetic Resources, Ministry of Education, Shanghai Ocean University, Shanghai, China.
Organizational Affiliation: