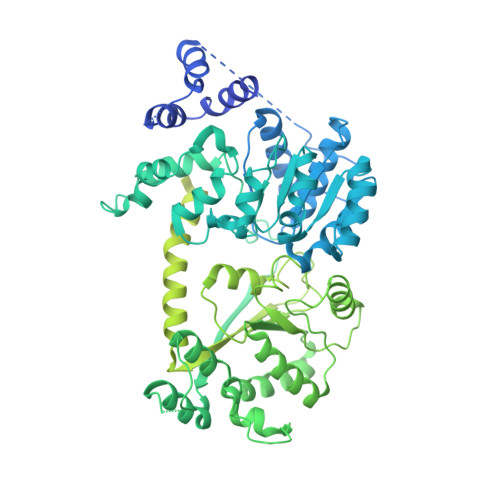
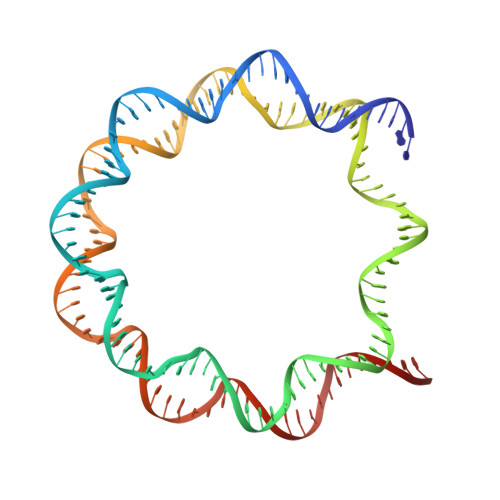
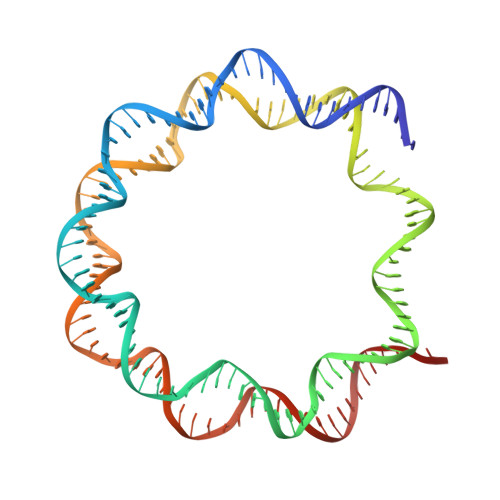
Structure of the ISW1a complex bound to the dinucleosome.
Li, L., Chen, K., Sia, Y., Hu, P., Ye, Y., Chen, Z.(2024) Nat Struct Mol Biol 31: 266-274
- PubMed: 38177688 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-023-01174-6
- Primary Citation Related Structures:
7X3T, 7X3V, 7X3W, 7X3X - PubMed Abstract:
Nucleosomes are basic repeating units of chromatin and form regularly spaced arrays in cells. Chromatin remodelers alter the positions of nucleosomes and are vital in regulating chromatin organization and gene expression. Here we report the cryo-EM structure of chromatin remodeler ISW1a complex from Saccharomyces cerevisiae bound to the dinucleosome. Each subunit of the complex recognizes a different nucleosome. The motor subunit binds to the mobile nucleosome and recognizes the acidic patch through two arginine residues, while the DNA-binding module interacts with the entry DNA at the nucleosome edge. This nucleosome-binding mode provides the structural basis for linker DNA sensing of the motor. Notably, the Ioc3 subunit recognizes the disk face of the adjacent nucleosome through interacting with the H4 tail, the acidic patch and the nucleosomal DNA, which plays a role in the spacing activity in vitro and in nucleosome organization and cell fitness in vivo. Together, these findings support the nucleosome spacing activity of ISW1a and add a new mode of nucleosome remodeling in the context of a chromatin environment.
- MOE Key Laboratory of Protein Science, Tsinghua University, Beijing, P.R. China.
Organizational Affiliation: