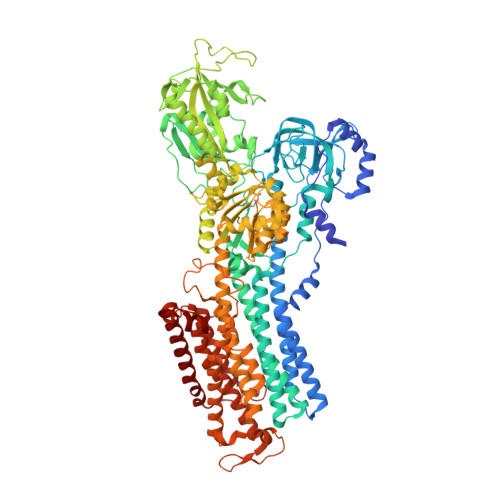
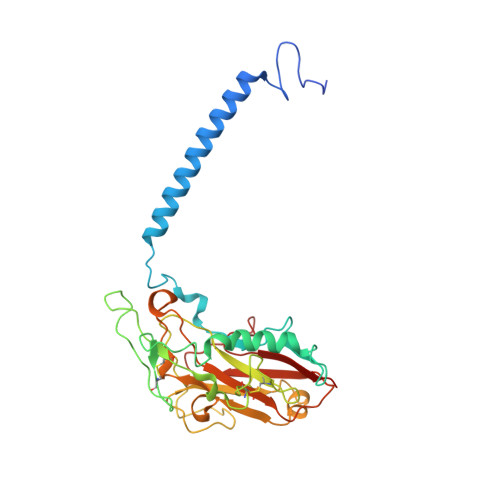
Cryoelectron microscopy of Na + ,K + -ATPase in the two E2P states with and without cardiotonic steroids.
Kanai, R., Cornelius, F., Vilsen, B., Toyoshima, C.(2022) Proc Natl Acad Sci U S A 119: e2123226119-e2123226119
- PubMed: 35380894 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2123226119
- Primary Citation Related Structures:
7WYS, 7WYT, 7WYU, 7WYV, 7WYW, 7WYX, 7WYY, 7WYZ, 7WZ0 - PubMed Abstract:
Cryoelectron microscopy (cryo-EM) was applied to Na+,K+-ATPase (NKA) to determine the structures of two E2P states, one (E2PATP) formed by ATP and Mg2+ in the forward reaction, and the other (E2PPi) formed by inorganic phosphate (Pi) and Mg2+ in the backward reaction, with and without ouabain or istaroxime, representatives of classical and new-generation cardiotonic steroids (CTSs). These two E2P states exhibit different biochemical properties. In particular, K+-sensitive acceleration of the dephosphorylation reaction is not observed with E2PPi, attributed to the presence of a Mg2+ ion in the transmembrane cation binding sites. The cryo-EM structures of NKA demonstrate that the two E2P structures are nearly identical but Mg2+ in the transmembrane binding cavity is identified only in E2PPi, corroborating the idea that it should be denoted as E2PPi·Mg2+. We can now explain why the absence of transmembrane Mg2+ in E2PATP confers the K+ sensitivity in dephosphorylation. In addition, we show that ATP bridges the actuator (A) and nucleotide binding (N) domains, stabilizing the E2PATP state; CTS binding causes hardly any changes in the structure of NKA, both in E2PATP and E2PPi·Mg2+, indicating that the binding mechanism is conformational selection; and istaroxime binds to NKA, extending its aminoalkyloxime group deep into the cation binding site. This orientation is upside down compared to that of classical CTSs with respect to the steroid ring. Notably, mobile parts of NKA are resolved substantially better in the electron microscopy (EM) maps than in previous X-ray structures, including sugars sticking out from the β-subunit and many phospholipid molecules.
- Institute for Quantitative Biosciences, The University of Tokyo, Bunkyo-ku, Tokyo 113-0032, Japan.
Organizational Affiliation: