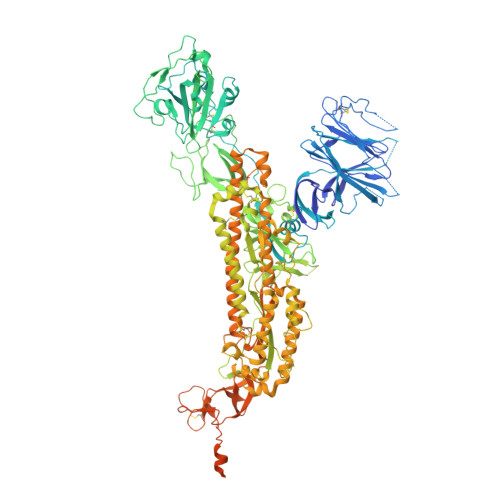
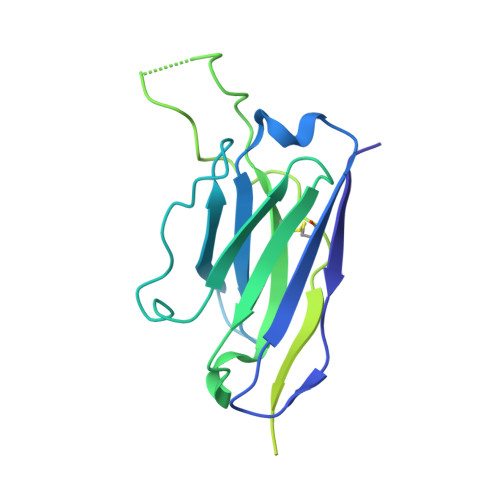
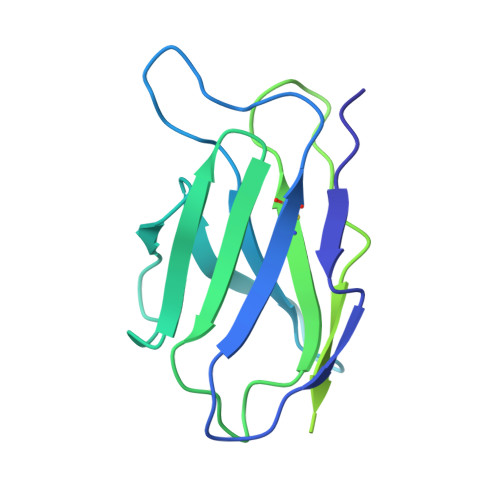
35B5 antibody potently neutralizes SARS-CoV-2 Omicron by disrupting the N-glycan switch via a conserved spike epitope.
Wang, X., Chen, X., Tan, J., Yue, S., Zhou, R., Xu, Y., Lin, Y., Yang, Y., Zhou, Y., Deng, K., Chen, Z., Ye, L., Zhu, Y.(2022) Cell Host Microbe 30: 887
- PubMed: 35436443 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.chom.2022.03.035
- Primary Citation Related Structures:
7WLY, 7WLZ - PubMed Abstract:
The SARS-CoV-2 Omicron variant harbors more than 30 mutations in the spike protein, leading to immune evasion from many therapeutic neutralizing antibodies. We reveal that a receptor-binding domain (RBD)-targeting monoclonal antibody, 35B5, exhibits potent neutralizing efficacy to Omicron. Cryo-electron microscopy structures of the extracellular domain trimer of Omicron spike with 35B5 Fab reveal that Omicron spike exhibits tight trimeric packing and high thermostability, as well as significant antigenic shifts and structural changes, within the RBD, N-terminal domain (NTD), and subdomains 1 and 2. However, these changes do not affect targeting of the invariant 35B5 epitope. 35B5 potently neutralizes SARS-CoV-2 Omicron and other variants by causing significant conformational changes within a conserved N-glycan switch that controls the transition of RBD from the "down" state to the "up" state, which allows recognition of the host entry receptor ACE2. This mode of action and potent neutralizing capacity of 35B5 indicate its potential therapeutic application for SARS-CoV-2.
- Department of Gastroenterology of the Second Affiliated Hospital, School of Medicine and Life Sciences Institute, Zhejiang University, Hangzhou 310058, China; MOE Laboratory for Biosystems Homeostasis and Protection, Life Sciences Institute, Zhejiang University, Hangzhou, Zhejiang 310058, China.
Organizational Affiliation: