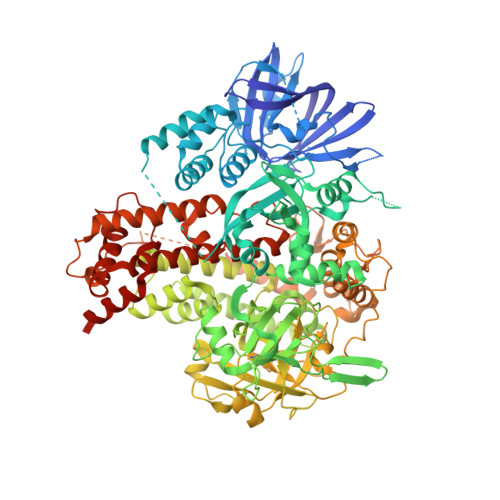
Structure of plant RNA-DEPENDENT RNA POLYMERASE 2, an enzyme involved in small interfering RNA production.
Du, X., Yang, Z., Ariza, A.J.F., Wang, Q., Xie, G., Li, S., Du, J.(2022) Plant Cell 34: 2140-2149
- PubMed: 35188193 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/plcell/koac067
- Primary Citation Related Structures:
7W82, 7W84, 7W88 - PubMed Abstract:
In plants, the biogenesis of small interfering RNA (siRNA) requires a family of RNA-dependent RNA polymerases that convert single-stranded RNA (ssRNA) into double-stranded RNA (dsRNA), which is subsequently cleaved into defined lengths by Dicer endonucleases. Here, we determined the structure of maize (Zea mays) RNA-DEPENDENT RNA POLYMERASE 2 (ZmRDR2) in the closed and open conformations. The core catalytic region of ZmRDR2 possesses the canonical DNA-dependent RNA polymerase (DdRP) catalytic sites, pointing to a shared RNA production mechanism between DdRPs and plant RDR-family proteins. Apo-ZmRDR2 adopts a highly compact structure, representing an inactive closed conformation. By contrast, adding RNA induced a significant conformational change in the ZmRDR2 Head domain that opened the RNA binding tunnel, suggesting this is an active elongation conformation of ZmRDR2. Overall, our structural studies trapped both the active and inactive conformations of ZmRDR2, providing insights into the molecular mechanism of dsRNA synthesis during plant siRNA production.
- Department of Biochemistry and Molecular Biology, International Cancer Center, Shenzhen University Health Science Center, Shenzhen 518060, China.
Organizational Affiliation: