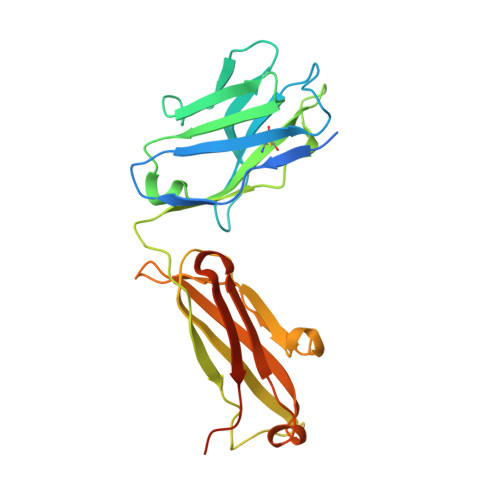
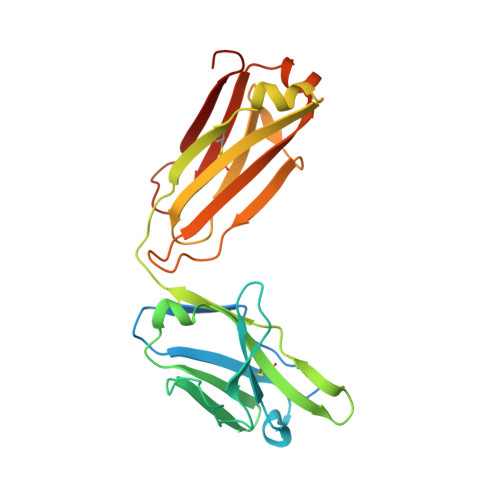
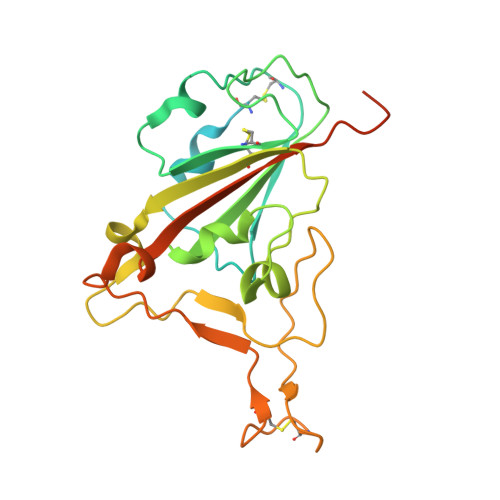
Computational design of a neutralizing antibody with picomolar binding affinity for all concerning SARS-CoV-2 variants.
Jeong, B.S., Cha, J.S., Hwang, I., Kim, U., Adolf-Bryfogle, J., Coventry, B., Cho, H.S., Kim, K.D., Oh, B.H.(2022) MAbs 14: 2021601-2021601
- PubMed: 35030983 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1080/19420862.2021.2021601
- Primary Citation Related Structures:
7VYR - PubMed Abstract:
Coronavirus disease 2019, caused by SARS-CoV-2, remains an on-going pandemic, partly due to the emergence of variant viruses that can "break-through" the protection of the current vaccines and neutralizing antibodies (nAbs), highlighting the needs for broadly nAbs and next-generation vaccines. We report an antibody that exhibits breadth and potency in binding the receptor-binding domain (RBD) of the virus spike glycoprotein across SARS coronaviruses. Initially, a lead antibody was computationally discovered and crystallographically validated that binds to a highly conserved surface of the RBD of wild-type SARS-CoV-2. Subsequently, through experimental affinity enhancement and computational affinity maturation, it was further developed to bind the RBD of all concerning SARS-CoV-2 variants, SARS-CoV-1 and pangolin coronavirus with pico-molar binding affinities, consistently exhibited strong neutralization activity against wild-type SARS-CoV-2 and the Alpha and Delta variants. These results identify a vulnerable target site on coronaviruses for development of pan-sarbecovirus nAbs and vaccines.
- Department of Biological Sciences, Kaist Institute for the Biocentury, Korea Advanced Institute of Science and Technology, Daejeon, Republic of Korea.
Organizational Affiliation: