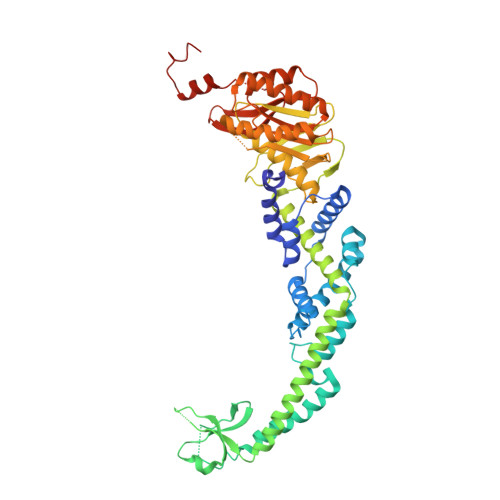
Structural and functional insights into the mechanism by which MutS2 recognizes a DNA junction.
Fukui, K., Inoue, M., Murakawa, T., Baba, S., Kumasaka, T., Yano, T.(2022) Structure 30: 973-982.e4
- PubMed: 35439431 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2022.03.014
- Primary Citation Related Structures:
7VUF, 7VUK - PubMed Abstract:
MutS family proteins are classified into MutS-I and -II lineages: MutS-I recognizes mismatched DNA and initiates mismatch repair, whereas MutS-II recognizes DNA junctions to modulate recombination. MutS-I forms dimeric clamp-like structures enclosing the mismatched DNA, and its composite ATPase sites regulate DNA-binding modes. Meanwhile, the structures of MutS-II have not been determined; accordingly, it remains unknown how MutS-II recognizes DNA junctions and how nucleotides control DNA binding. Here, we solved the ligand-free and ADP-bound crystal structures of bacterial MutS2 belonging to MutS-II. MutS2 also formed a dimeric clamp-like structure with composite ATPase sites. The ADP-bound MutS2 was more flexible compared to the ligand-free form and could be more suitable for DNA entry. The inner hole of the MutS2 clamp was two times larger than that of MutS-I, and site-directed mutagenesis analyses revealed DNA-binding sites at the inner hole. Based on these, a model is proposed that describes how MutS2 recognizes DNA junctions.
- Department of Biochemistry, Faculty of Medicine, Osaka Medical and Pharmaceutical University, Takatsuki, Osaka 569-8686, Japan. Electronic address: kenji.fukui@ompu.ac.jp.
Organizational Affiliation: