A distinctive ligand recognition mechanism by the human vasoactive intestinal polypeptide receptor 2.
Xu, Y., Feng, W., Zhou, Q., Liang, A., Li, J., Dai, A., Zhao, F., Yan, J., Chen, C.W., Li, H., Zhao, L.H., Xia, T., Jiang, Y., Xu, H.E., Yang, D., Wang, M.W.(2022) Nat Commun 13: 2272-2272
- PubMed: 35477937 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-30041-z
- Primary Citation Related Structures:
7VQX, 7WBJ - PubMed Abstract:
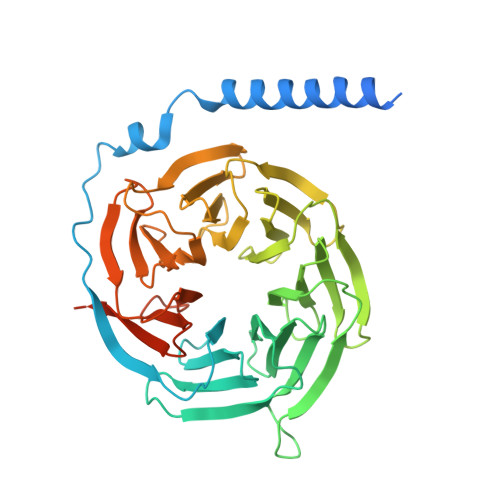
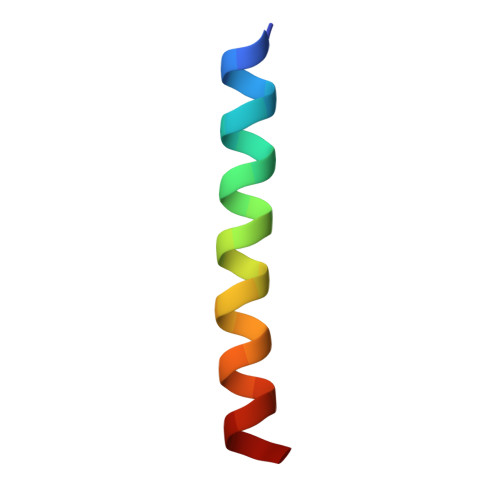
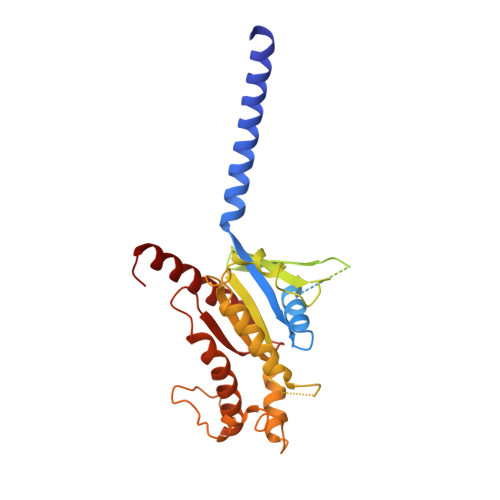
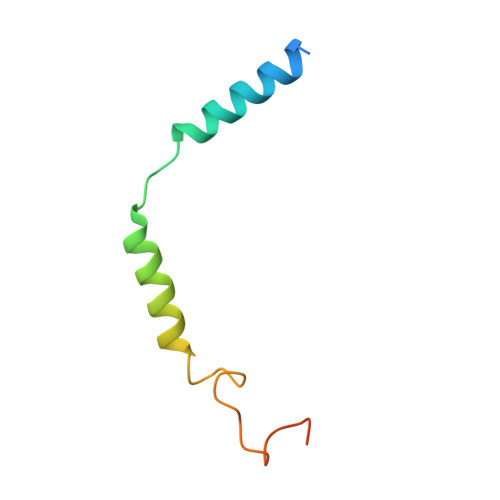
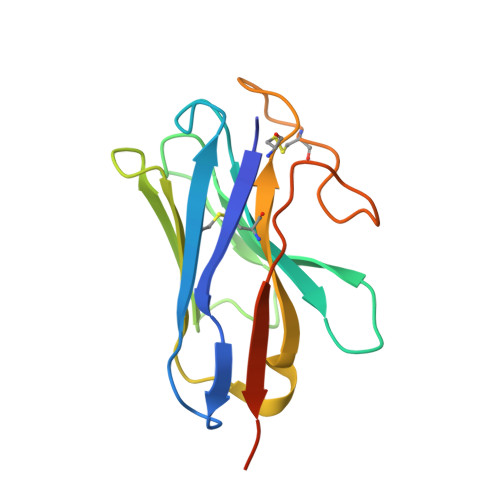
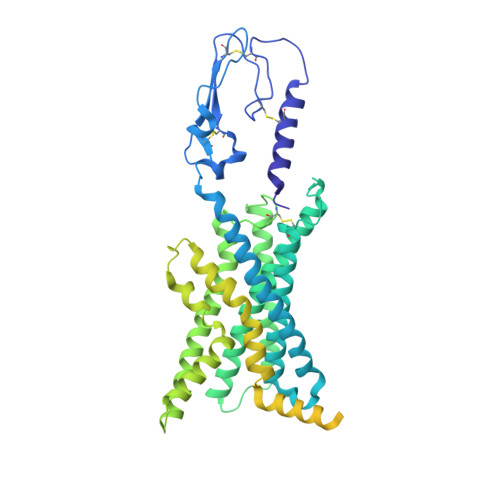
Class B1 of G protein-coupled receptors (GPCRs) comprises 15 members activated by physiologically important peptide hormones. Among them, vasoactive intestinal polypeptide receptor 2 (VIP2R) is expressed in the central and peripheral nervous systems and involved in a number of pathophysiological conditions, including pulmonary arterial hypertension, autoimmune and psychiatric disorders, in which it is thus a valuable drug target. Here, we report the cryo-electron microscopy structure of the human VIP2R bound to its endogenous ligand PACAP27 and the stimulatory G protein. Different from all reported peptide-bound class B1 GPCR structures, the N-terminal α-helix of VIP2R adopts a unique conformation that deeply inserts into a cleft between PACAP27 and the extracellular loop 1, thereby stabilizing the peptide-receptor interface. Its truncation or extension significantly decreased VIP2R-mediated cAMP accumulation. Our results provide additional information on peptide recognition and receptor activation among class B1 GPCRs and may facilitate the design of better therapeutics.
- Department of Pharmacology, School of Basic Medical Sciences, Fudan University, Shanghai, 200032, China.
Organizational Affiliation: