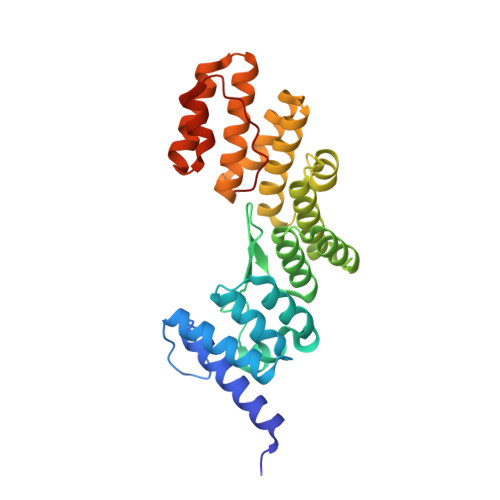
The crystal structure of the TPR domain of the EccA5 ATPase and demonstration of its interaction with EspG5 from the mycobacterial ESX-5 pathway.
Sharma, V.K., Vishwakarma, J., Kabrambam, R., Kumar, S., Ramachandran, R.(2026) FEBS Lett
- PubMed: 41761896 Search on PubMed
- DOI: https://doi.org/10.1002/1873-3468.70315
- Primary Citation Related Structures:
7VEP - PubMed Abstract:
The ESX-5 secretion system in Mycobacterium tuberculosis exports PE/PPE virulence factors, with EccA5, an AAA+ ATPase, playing a pivotal role. We solved the crystal structure of EccA5's N-terminal TPR domain (EccA5NT) at 2.15 Å, revealing a monomeric fold with six TPR motifs and a variable β-finger. Biophysical studies, including SAXS and size exclusion chromatography, confirm its monomeric state. A flexible loop (residues 137-148) suggests dynamic substrate interactions. SPR, SAXS and in silico docking show moderate binding (K D = 3.43 μm) between EccA5NT's β-finger and EspG5's β2-β3 loop, indicating a role in PE/PPE-EspG5 complex disassembly. These findings elucidate the role of EccA5 in ESX-5-mediated secretion.
- Biochemistry and Structural Biology Division, CSIR-Central Drug Research Institute, Lucknow, India.
Organizational Affiliation: