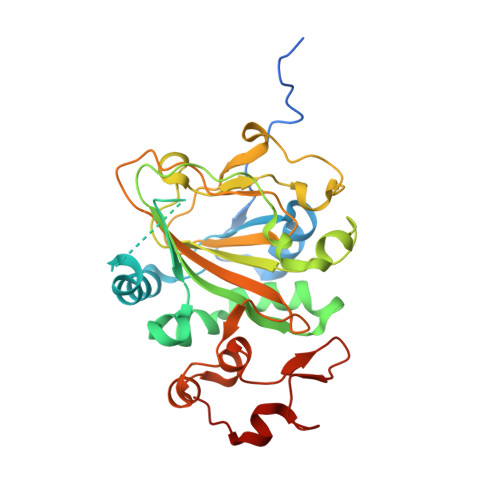
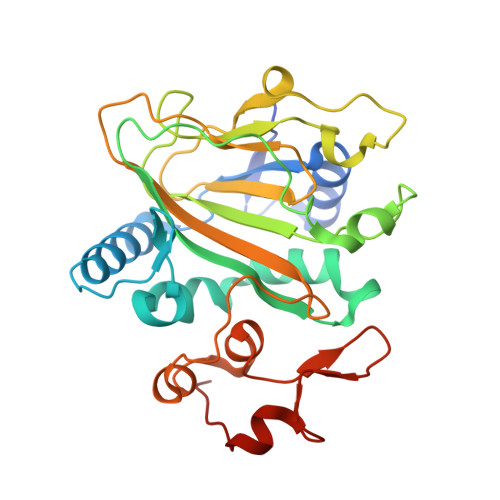
Heterodimeric Non-heme Iron Enzymes in Fungal Meroterpenoid Biosynthesis.
Li, X., Awakawa, T., Mori, T., Ling, M., Hu, D., Wu, B., Abe, I.(2021) J Am Chem Soc 143: 21425-21432
- PubMed: 34881885 Search on PubMed
- DOI: https://doi.org/10.1021/jacs.1c11548
- Primary Citation Related Structures:
7VBQ, 7VBR - PubMed Abstract:
Talaromyolides ( 1 - 6 ) are a group of unusual 6/6/6/6/6/6 hexacyclic meroterpenoids with (3 R )-6-hydroxymellein and 4,5-seco-drimane substructures, isolated from the marine fungus Talaromyces purpureogenus . We have identified the biosynthetic gene cluster tlxA-J by heterologous expression in Aspergillus , in vitro enzyme assays, and CRISPR-Cas9-based gene inactivation. Remarkably, the heterodimer of non-heme iron (NHI) enzymes, TlxJ-TlxI, catalyzes three steps of oxidation including a key reaction, hydroxylation at C-5 and C-9 of 12 , the intermediate with 3-ketohydroxydrimane scaffold, to facilitate a retro-aldol reaction, leading to the construction of the 4,5-secodrimane skeleton and characteristic ketal scaffold of 1 - 6 . The products of TlxJ-TlxI, 1 and 4 , were further hydroxylated at C-4'β by another NHI heterodimer, TlxA-TlxC, and acetylated by TlxB to yield the final products, 3 and 6 . The X-ray structural analysis coupled with site-directed mutagenesis provided insights into the heterodimer TlxJ-TlxI formation and its catalysis. This is the first report to show that two NHI proteins form a heterodimer for catalysis and utilizes a novel methodology to create functional oxygenase structures in secondary metabolite biosynthesis.
- Graduate School of Pharmaceutical Sciences, The University of Tokyo, Bunkyo-ku, Tokyo 113-0033, Japan.
Organizational Affiliation: