MERS S ectodomain trimer in complex with neutralizing antibody 6516
Wang, X., Zhao, J., Wang, Z., Wang, Y., Zeng, J.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Spike glycoprotein | 1,290 | Human betacoronavirus 2c EMC/2012 | Mutation(s): 0 | 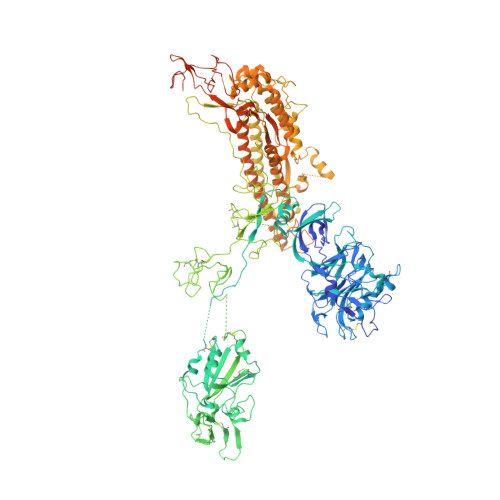 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | K0BRG7 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| antibody H | 226 | Homo sapiens | Mutation(s): 0 | 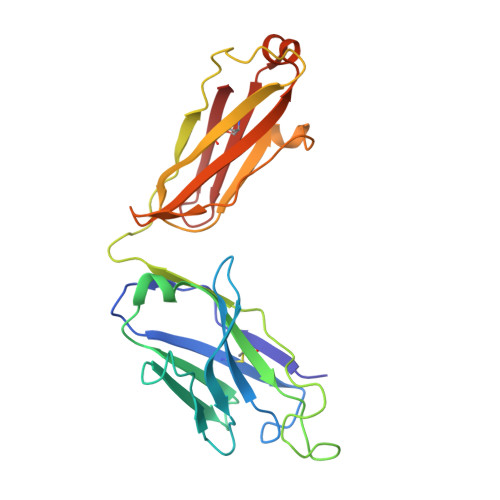 | |
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| antibody L | 215 | Homo sapiens | Mutation(s): 0 | 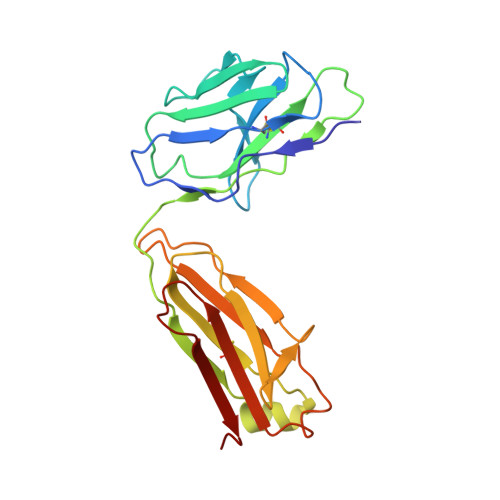 | |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Not funded | -- |