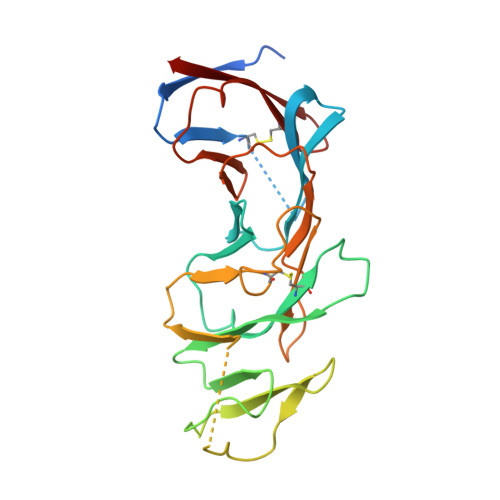
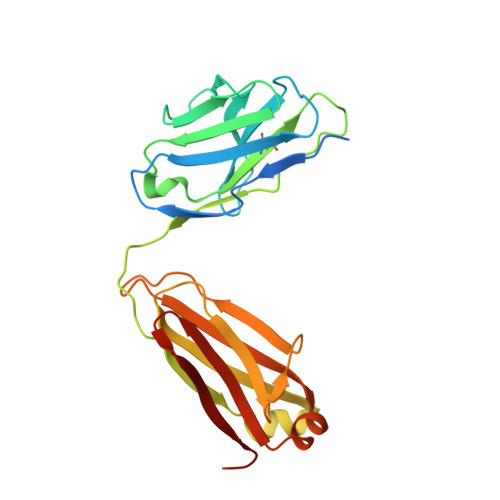
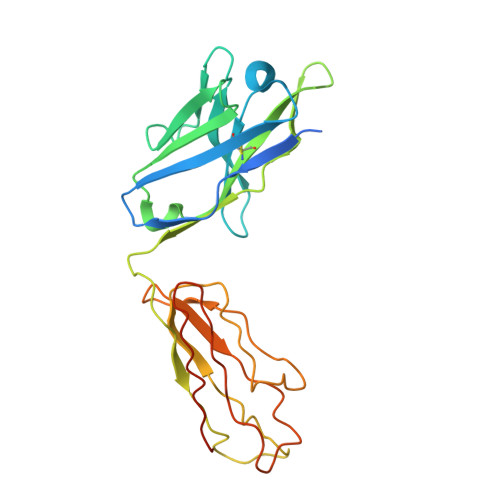
CD19 CAR antigen engagement mechanisms and affinity tuning.
He, C., Mansilla-Soto, J., Khanra, N., Hamieh, M., Bustos, V., Paquette, A.J., Garcia Angus, A., Shore, D.M., Rice, W.J., Khelashvili, G., Sadelain, M., Meyerson, J.R.(2023) Sci Immunol 8: eadf1426-eadf1426
- PubMed: 36867678 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciimmunol.adf1426
- Primary Citation Related Structures:
7URV, 7URX - PubMed Abstract:
Chimeric antigen receptor (CAR) T cell therapy relies on T cells that are guided by synthetic receptors to target and lyse cancer cells. CARs bind to cell surface antigens through an scFv (binder), the affinity of which is central to determining CAR T cell function and therapeutic success. CAR T cells targeting CD19 were the first to achieve marked clinical responses in patients with relapsed/refractory B cell malignancies and to be approved by the U.S. Food and Drug Administration (FDA). We report cryo-EM structures of CD19 antigen with the binder FMC63, which is used in four FDA-approved CAR T cell therapies (Kymriah, Yescarta, Tecartus, and Breyanzi), and the binder SJ25C1, which has also been used extensively in multiple clinical trials. We used these structures for molecular dynamics simulations, which guided creation of lower- or higher-affinity binders, and ultimately produced CAR T cells endowed with distinct tumor recognition sensitivities. The CAR T cells exhibited different antigen density requirements to trigger cytolysis and differed in their propensity to prompt trogocytosis upon contacting tumor cells. Our work shows how structural information can be applied to tune CAR T cell performance to specific target antigen densities.
- Department of Physiology and Biophysics, Weill Cornell Medical College, New York, NY, USA.
Organizational Affiliation: